Posterior Sampling Distribution for the Elements of the Matrix of Lagged Coefficients Over a Specific Time Interval or a Range of Time Intervals
Source:R/cTMed-posterior-beta.R
PosteriorBeta.RdThis function generates a posterior sampling distribution for the elements of the matrix of lagged coefficients \(\boldsymbol{\beta}\) over a specific time interval \(\Delta t\) or a range of time intervals using the first-order stochastic differential equation model drift matrix \(\boldsymbol{\Phi}\).
Arguments
- phi
List of numeric matrices. Each element of the list is a sample from the posterior distribution of the drift matrix (\(\boldsymbol{\Phi}\)). Each matrix should have row and column names pertaining to the variables in the system.
- delta_t
Numeric. Time interval (\(\Delta t\)).
- ncores
Positive integer. Number of cores to use. If
ncores = NULL, use a single core. Consider using multiple cores when number of replicationsRis a large value.- tol
Numeric. Smallest possible time interval to allow.
Value
Returns an object
of class ctmedmc which is a list with the following elements:
- call
Function call.
- args
Function arguments.
- fun
Function used ("PosteriorBeta").
- output
A list the length of which is equal to the length of
delta_t.
Each element in the output list has the following elements:
- est
A vector of total, direct, and indirect effects.
- thetahatstar
A matrix of Monte Carlo total, direct, and indirect effects.
Details
See Total().
References
Bollen, K. A. (1987). Total, direct, and indirect effects in structural equation models. Sociological Methodology, 17, 37. doi:10.2307/271028
Deboeck, P. R., & Preacher, K. J. (2015). No need to be discrete: A method for continuous time mediation analysis. Structural Equation Modeling: A Multidisciplinary Journal, 23 (1), 61-75. doi:10.1080/10705511.2014.973960
Pesigan, I. J. A., Russell, M. A., & Chow, S.-M. (2025). Inferences and effect sizes for direct, indirect, and total effects in continuous-time mediation models. Psychological Methods. doi:10.1037/met0000779
Ryan, O., & Hamaker, E. L. (2021). Time to intervene: A continuous-time approach to network analysis and centrality. Psychometrika, 87 (1), 214-252. doi:10.1007/s11336-021-09767-0
See also
Other Continuous-Time Mediation Functions:
BootBeta(),
BootBetaStd(),
BootDirectCentral(),
BootIndirectCentral(),
BootMed(),
BootMedStd(),
BootTotalCentral(),
DeltaBeta(),
DeltaBetaStd(),
DeltaDirectCentral(),
DeltaIndirectCentral(),
DeltaMed(),
DeltaMedStd(),
DeltaTotalCentral(),
Direct(),
DirectCentral(),
DirectStd(),
Indirect(),
IndirectCentral(),
IndirectStd(),
MCBeta(),
MCBetaStd(),
MCDirectCentral(),
MCIndirectCentral(),
MCMed(),
MCMedStd(),
MCPhi(),
MCPhiSigma(),
MCTotalCentral(),
Med(),
MedStd(),
PosteriorDirectCentral(),
PosteriorIndirectCentral(),
PosteriorMed(),
PosteriorTotalCentral(),
Total(),
TotalCentral(),
TotalStd(),
Trajectory()
Examples
phi <- matrix(
data = c(
-0.357, 0.771, -0.450,
0.0, -0.511, 0.729,
0, 0, -0.693
),
nrow = 3
)
colnames(phi) <- rownames(phi) <- c("x", "m", "y")
vcov_phi_vec <- matrix(
data = c(
0.00843, 0.00040, -0.00151,
-0.00600, -0.00033, 0.00110,
0.00324, 0.00020, -0.00061,
0.00040, 0.00374, 0.00016,
-0.00022, -0.00273, -0.00016,
0.00009, 0.00150, 0.00012,
-0.00151, 0.00016, 0.00389,
0.00103, -0.00007, -0.00283,
-0.00050, 0.00000, 0.00156,
-0.00600, -0.00022, 0.00103,
0.00644, 0.00031, -0.00119,
-0.00374, -0.00021, 0.00070,
-0.00033, -0.00273, -0.00007,
0.00031, 0.00287, 0.00013,
-0.00014, -0.00170, -0.00012,
0.00110, -0.00016, -0.00283,
-0.00119, 0.00013, 0.00297,
0.00063, -0.00004, -0.00177,
0.00324, 0.00009, -0.00050,
-0.00374, -0.00014, 0.00063,
0.00495, 0.00024, -0.00093,
0.00020, 0.00150, 0.00000,
-0.00021, -0.00170, -0.00004,
0.00024, 0.00214, 0.00012,
-0.00061, 0.00012, 0.00156,
0.00070, -0.00012, -0.00177,
-0.00093, 0.00012, 0.00223
),
nrow = 9
)
phi <- MCPhi(
phi = phi,
vcov_phi_vec = vcov_phi_vec,
R = 1000L
)$output
# Specific time interval ----------------------------------------------------
PosteriorBeta(
phi = phi,
delta_t = 1
)
#> Call:
#> PosteriorBeta(phi = phi, delta_t = 1)
#>
#> Total, Direct, and Indirect Effects
#>
#> effect interval est se R 2.5% 97.5%
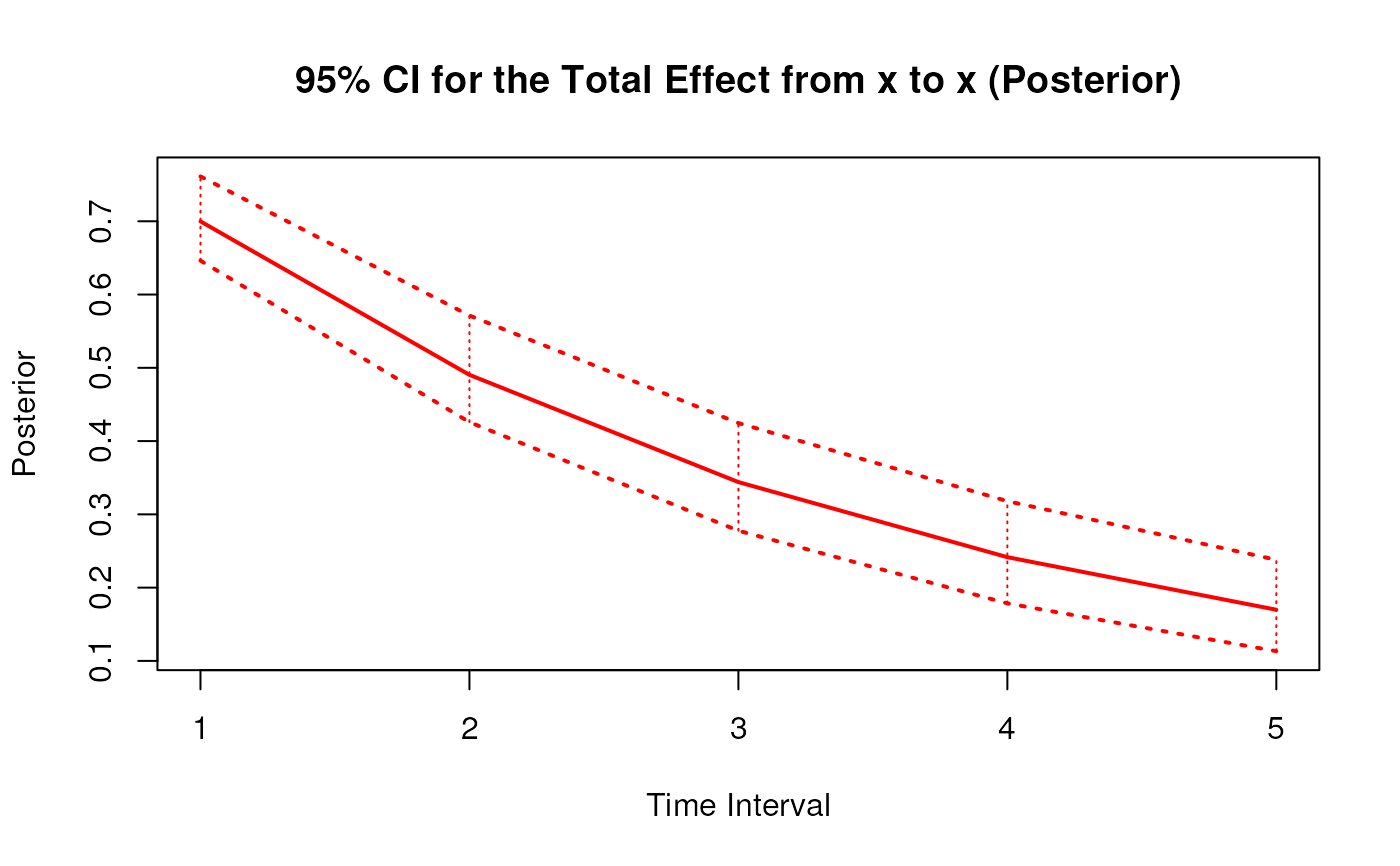
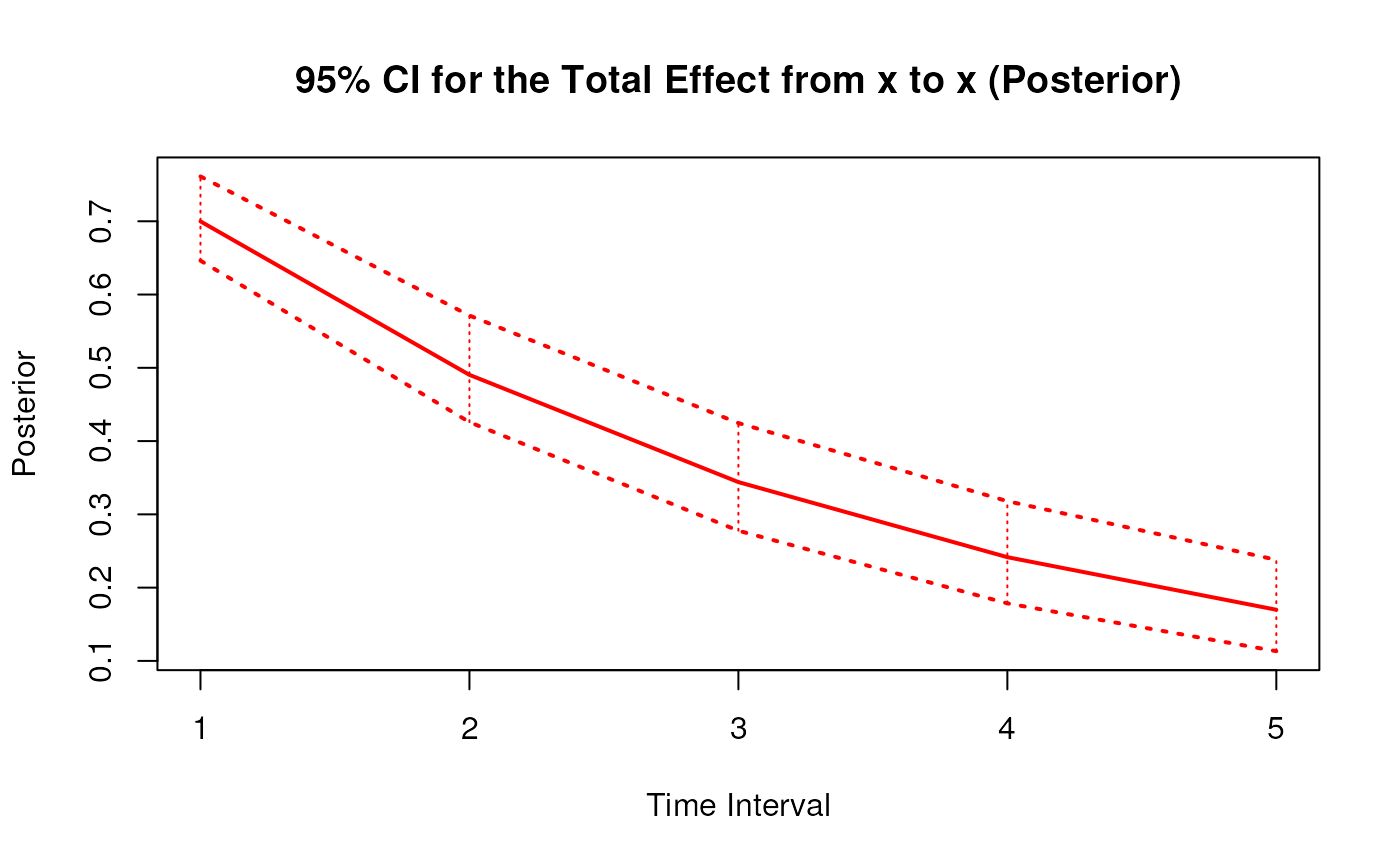
#> 1 from x to x 1 0.6998 0.0463 1000 0.6151 0.7991
#> 2 from x to m 1 0.4977 0.0345 1000 0.4344 0.5679
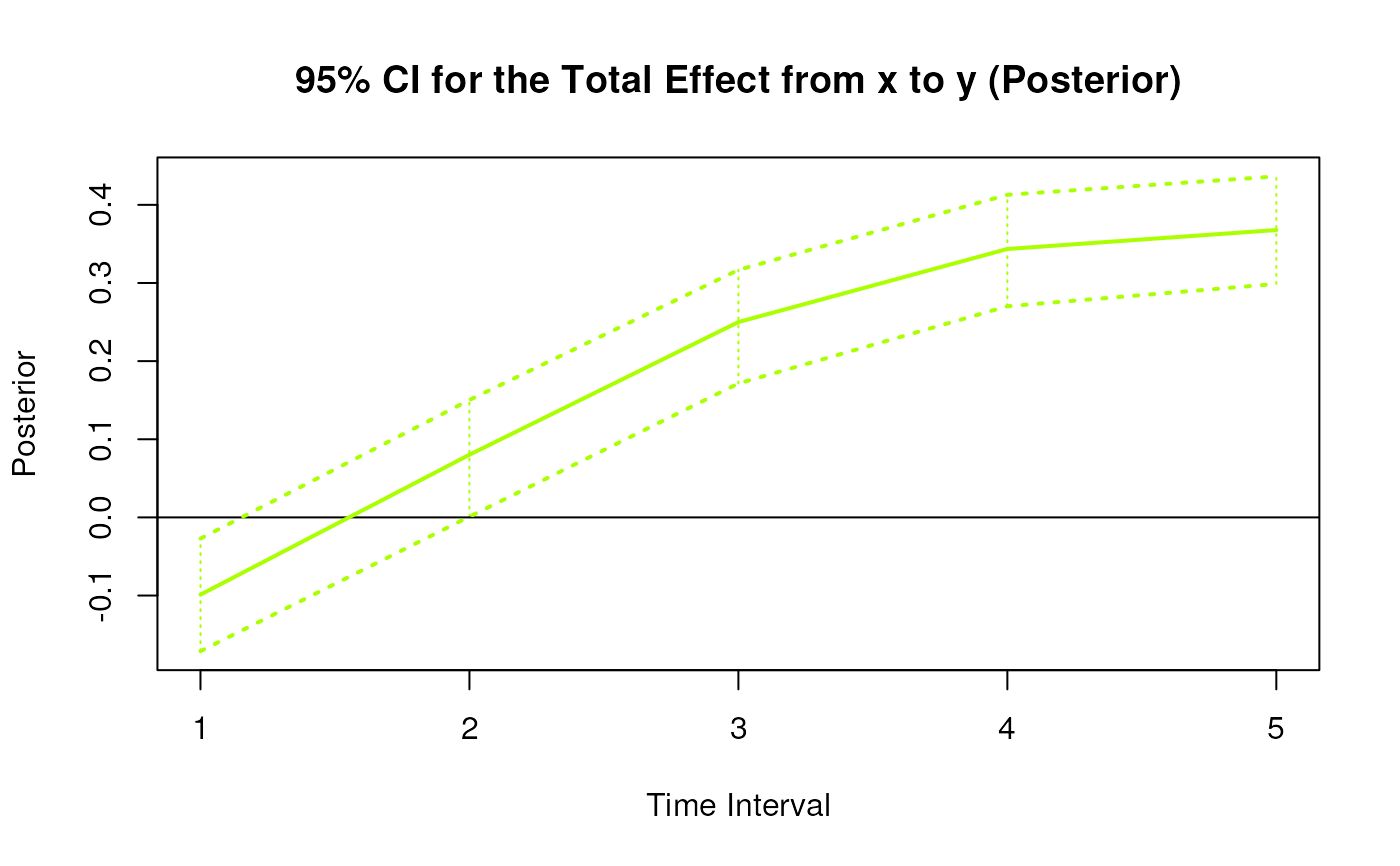
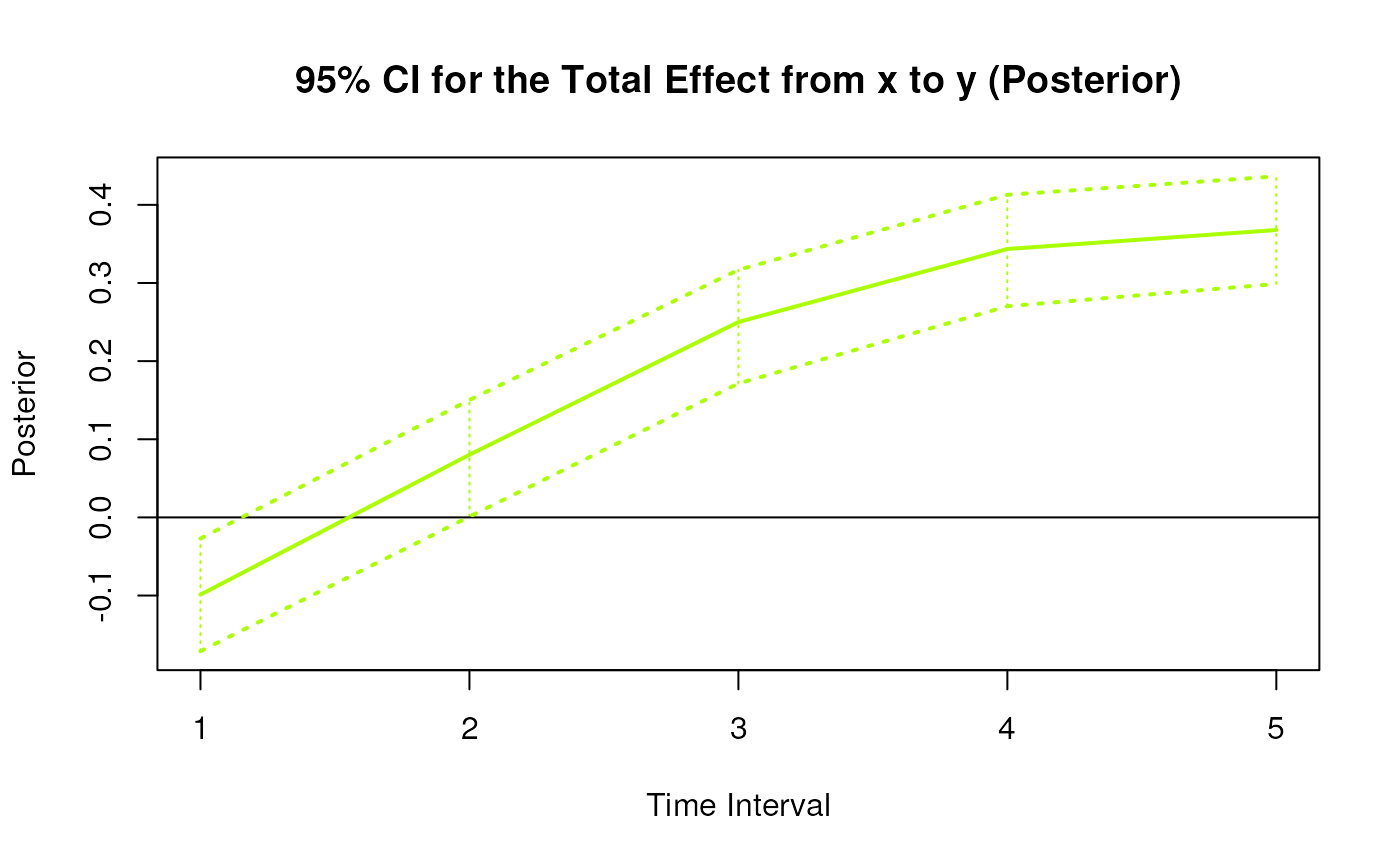
#> 3 from x to y 1 -0.1005 0.0307 1000 -0.1602 -0.0410
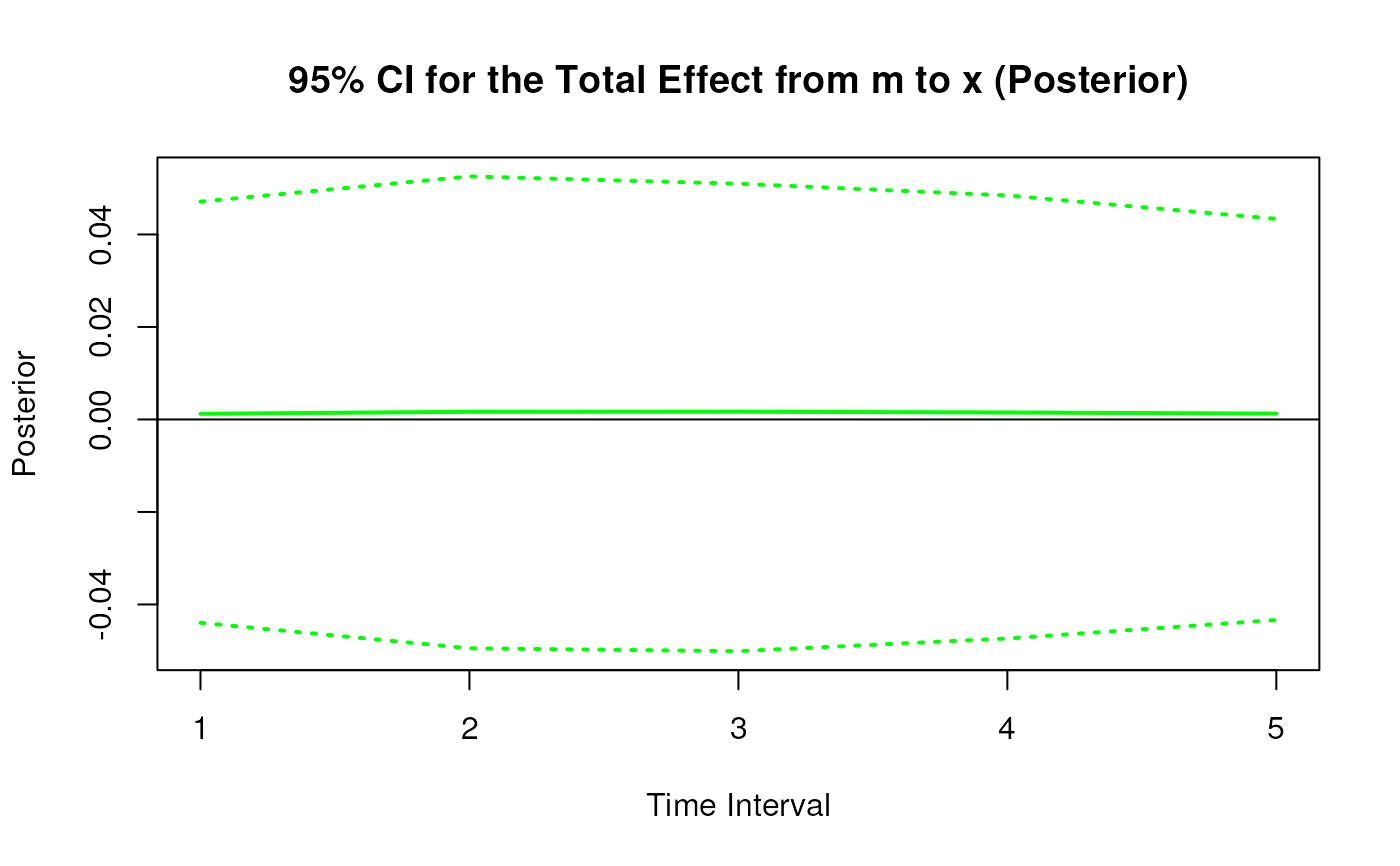
#> 4 from m to x 1 0.0021 0.0439 1000 -0.0829 0.0894
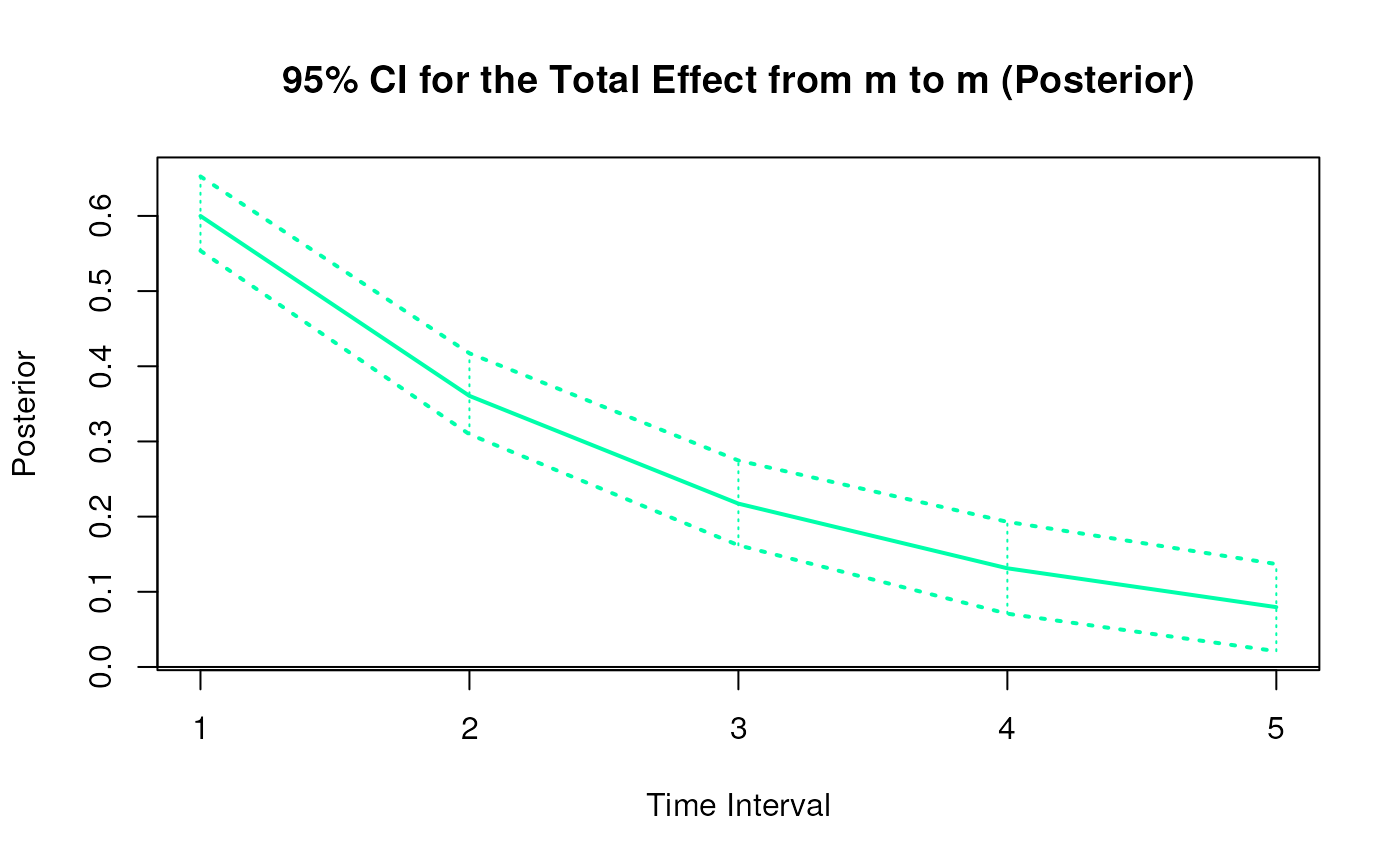
#> 5 from m to m 1 0.6013 0.0322 1000 0.5393 0.6640
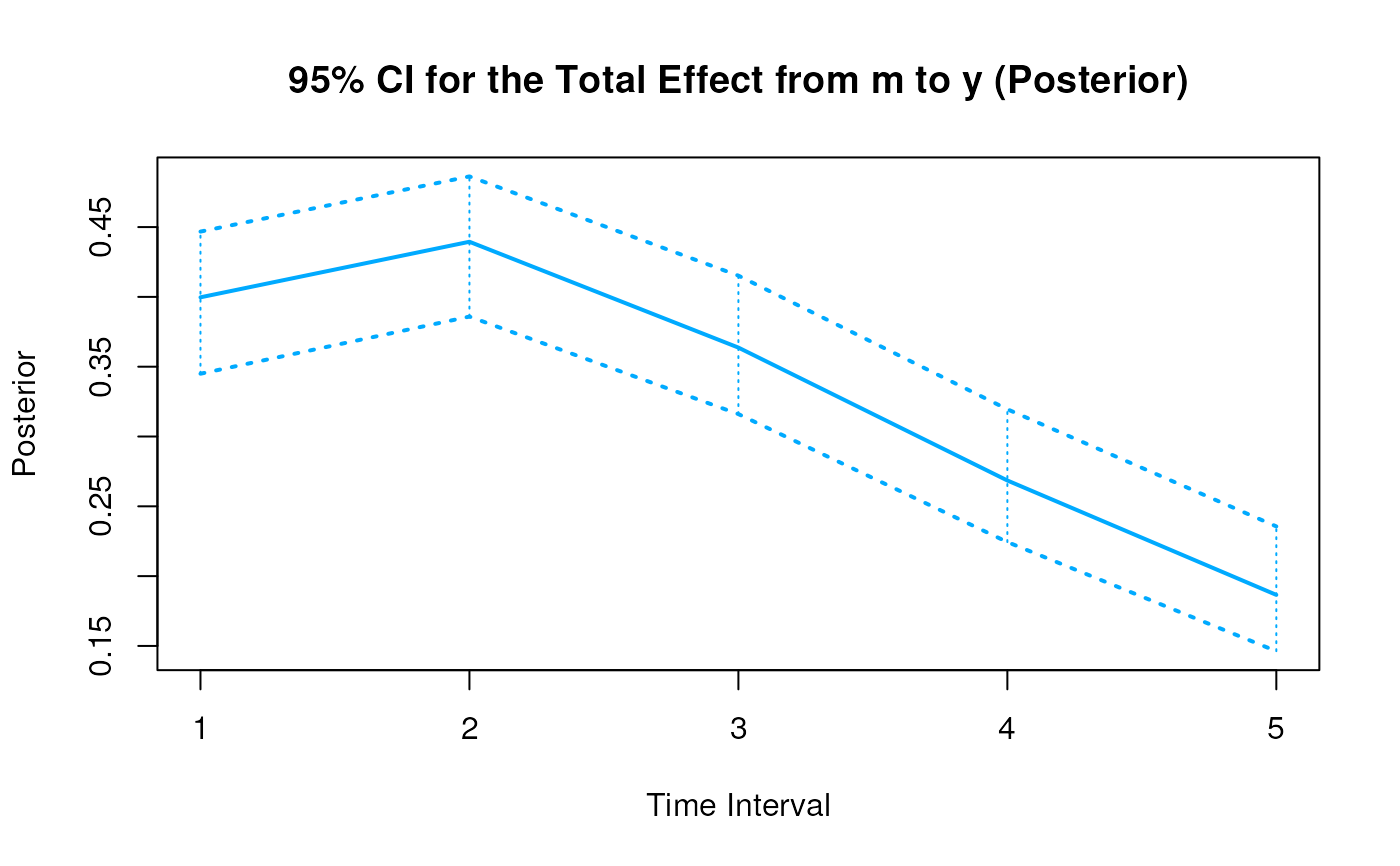
#> 6 from m to y 1 0.3994 0.0283 1000 0.3447 0.4551
#> 7 from y to x 1 0.0009 0.0427 1000 -0.0843 0.0824
#> 8 from y to m 1 0.0000 0.0307 1000 -0.0589 0.0598
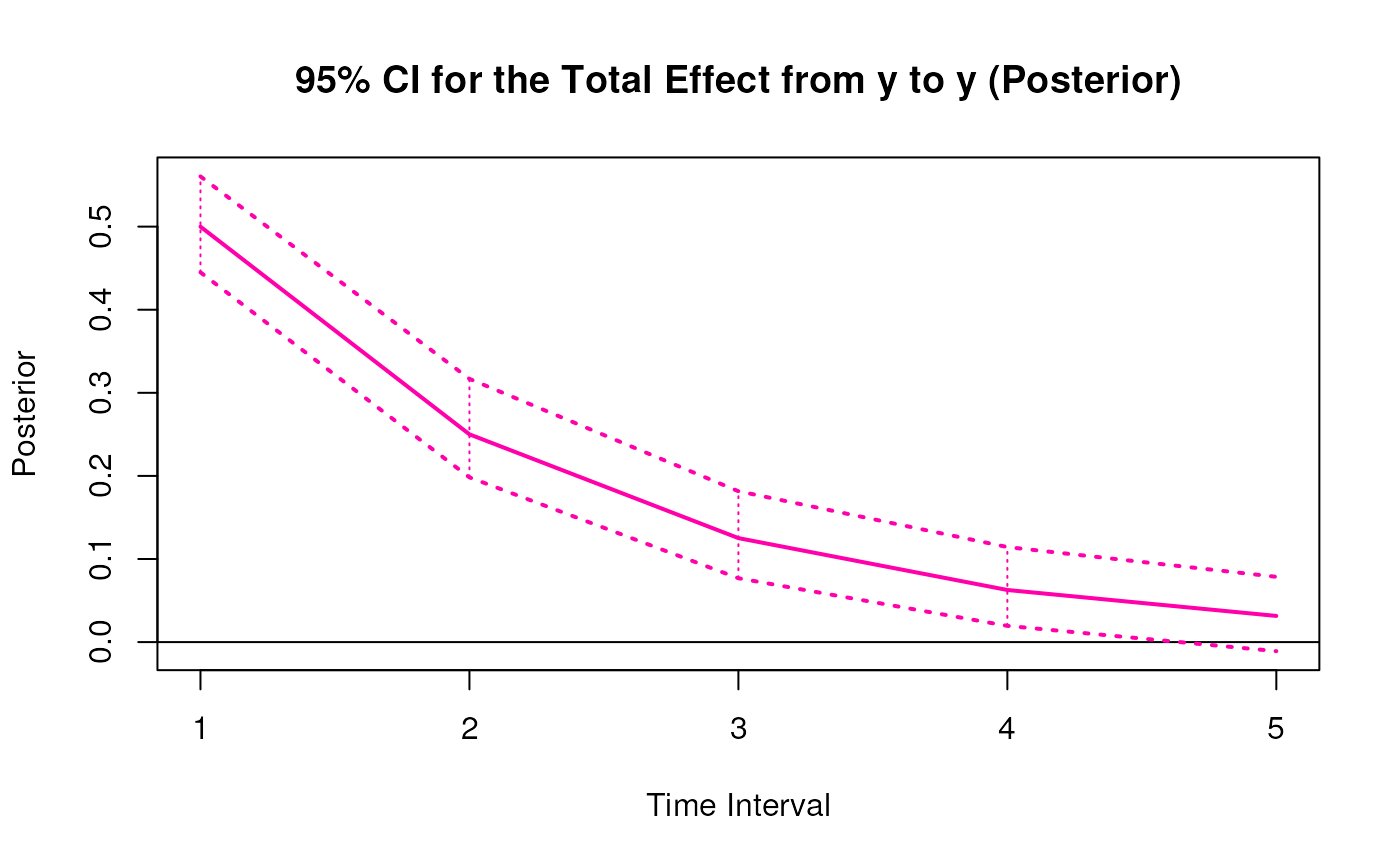
#> 9 from y to y 1 0.4997 0.0265 1000 0.4500 0.5523
# Range of time intervals ---------------------------------------------------
posterior <- PosteriorBeta(
phi = phi,
delta_t = 1:5
)
plot(posterior)


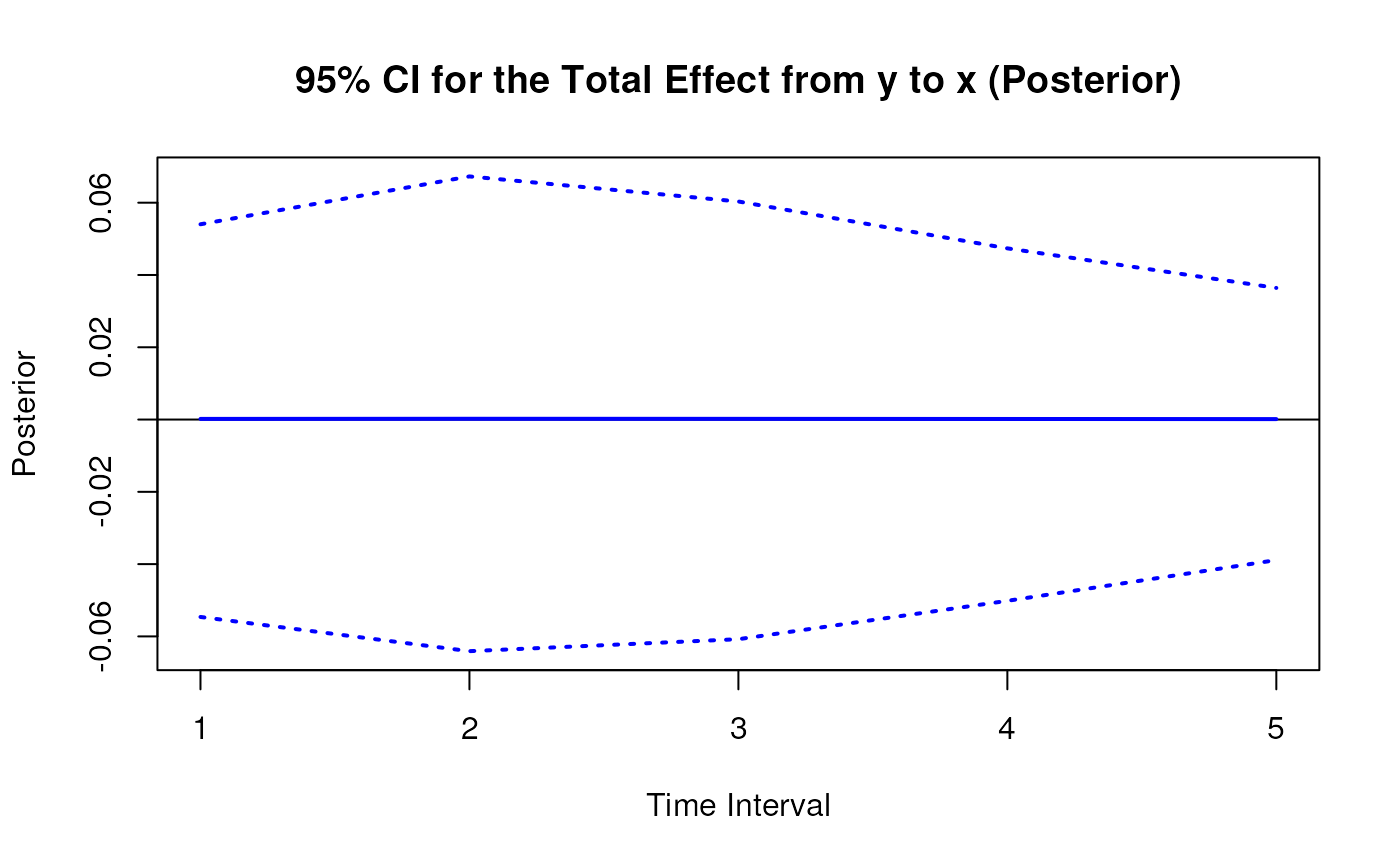

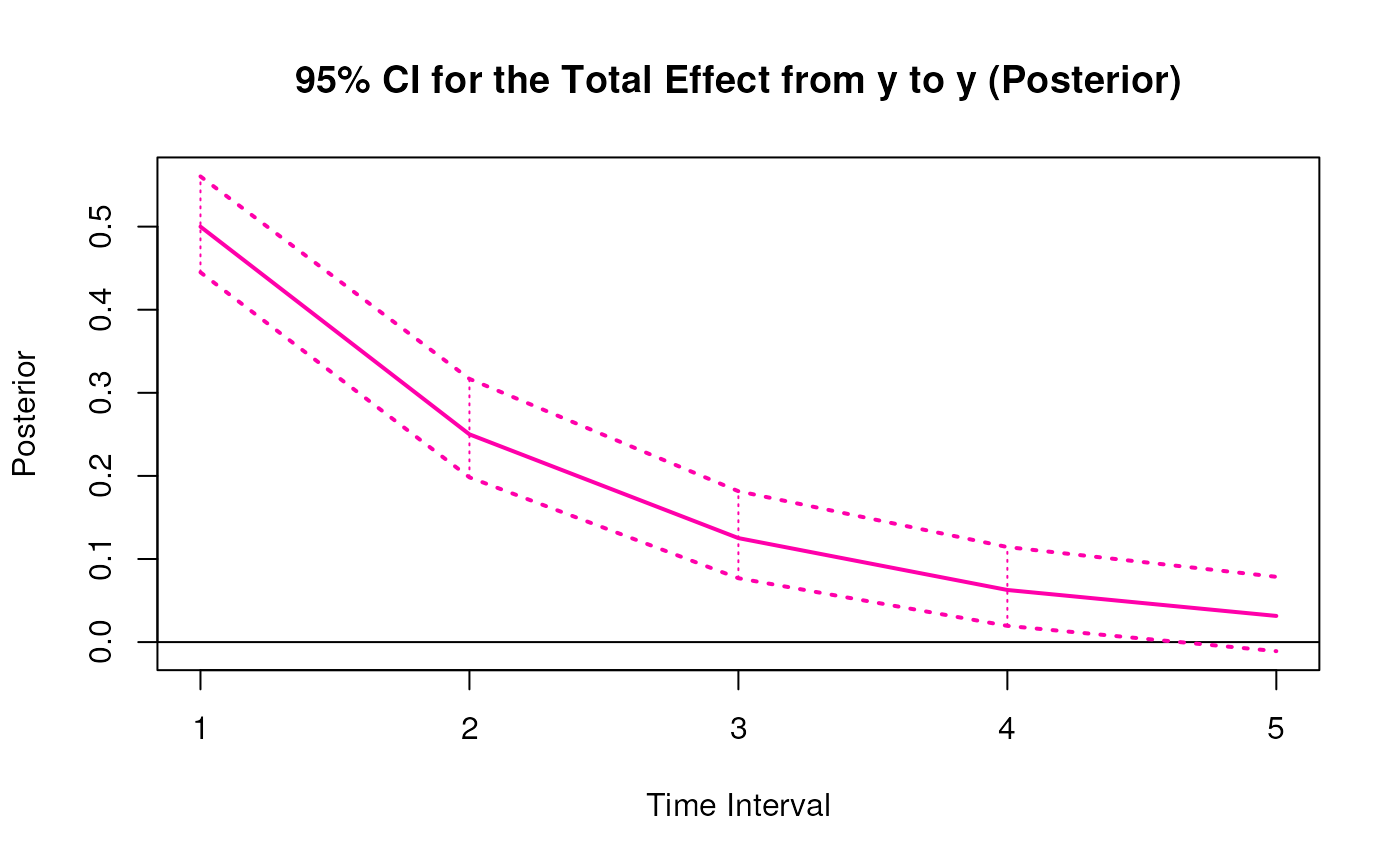 # Methods -------------------------------------------------------------------
# PosteriorBeta has a number of methods including
# print, summary, confint, and plot
print(posterior)
#> Call:
#> PosteriorBeta(phi = phi, delta_t = 1:5)
#>
#> Total, Direct, and Indirect Effects
#>
#> effect interval est se R 2.5% 97.5%
#> 1 from x to x 1 0.6998 0.0463 1000 0.6151 0.7991
#> 2 from x to m 1 0.4977 0.0345 1000 0.4344 0.5679
#> 3 from x to y 1 -0.1005 0.0307 1000 -0.1602 -0.0410
#> 4 from m to x 1 0.0021 0.0439 1000 -0.0829 0.0894
#> 5 from m to m 1 0.6013 0.0322 1000 0.5393 0.6640
#> 6 from m to y 1 0.3994 0.0283 1000 0.3447 0.4551
#> 7 from y to x 1 0.0009 0.0427 1000 -0.0843 0.0824
#> 8 from y to m 1 0.0000 0.0307 1000 -0.0589 0.0598
#> 9 from y to y 1 0.4997 0.0265 1000 0.4500 0.5523
#> 10 from x to x 2 0.4907 0.0543 1000 0.3994 0.6117
#> 11 from x to m 2 0.6476 0.0525 1000 0.5522 0.7579
#> 12 from x to y 2 0.0783 0.0347 1000 0.0050 0.1411
#> 13 from m to x 2 0.0031 0.0515 1000 -0.0970 0.1046
#> 14 from m to m 2 0.3626 0.0493 1000 0.2690 0.4606
#> 15 from m to y 2 0.4395 0.0332 1000 0.3789 0.5078
#> 16 from y to x 2 0.0011 0.0516 1000 -0.1025 0.0999
#> 17 from y to m 2 0.0004 0.0497 1000 -0.1008 0.0965
#> 18 from y to y 2 0.2496 0.0306 1000 0.1939 0.3104
#> 19 from x to x 3 0.3448 0.0547 1000 0.2528 0.4631
#> 20 from x to m 3 0.6336 0.0647 1000 0.5207 0.7680
#> 21 from x to y 3 0.2484 0.0356 1000 0.1777 0.3183
#> 22 from m to x 3 0.0034 0.0500 1000 -0.0930 0.1029
#> 23 from m to m 3 0.2195 0.0594 1000 0.1071 0.3407
#> 24 from m to y 3 0.3641 0.0335 1000 0.3028 0.4321
#> 25 from y to x 3 0.0010 0.0473 1000 -0.0955 0.0913
#> 26 from y to m 3 0.0008 0.0598 1000 -0.1181 0.1113
#> 27 from y to y 3 0.1248 0.0288 1000 0.0711 0.1838
#> 28 from x to x 4 0.2428 0.0539 1000 0.1526 0.3541
#> 29 from x to m 4 0.5526 0.0716 1000 0.4337 0.7058
#> 30 from x to y 4 0.3425 0.0398 1000 0.2701 0.4224
#> 31 from m to x 4 0.0031 0.0461 1000 -0.0876 0.0980
#> 32 from m to m 4 0.1337 0.0636 1000 0.0143 0.2657
#> 33 from m to y 4 0.2693 0.0351 1000 0.2060 0.3392
#> 34 from y to x 4 0.0008 0.0391 1000 -0.0804 0.0731
#> 35 from y to m 4 0.0009 0.0617 1000 -0.1224 0.1118
#> 36 from y to y 4 0.0626 0.0304 1000 0.0032 0.1251
#> 37 from x to x 5 0.1714 0.0531 1000 0.0838 0.2841
#> 38 from x to m 5 0.4531 0.0749 1000 0.3274 0.6108
#> 39 from x to y 5 0.3674 0.0449 1000 0.2855 0.4639
#> 40 from m to x 5 0.0027 0.0411 1000 -0.0776 0.0890
#> 41 from m to m 5 0.0819 0.0635 1000 -0.0414 0.2158
#> 42 from m to y 5 0.1876 0.0374 1000 0.1198 0.2662
#> 43 from y to x 5 0.0006 0.0307 1000 -0.0638 0.0584
#> 44 from y to m 5 0.0010 0.0577 1000 -0.1192 0.1073
#> 45 from y to y 5 0.0316 0.0342 1000 -0.0349 0.0988
summary(posterior)
#> Call:
#> PosteriorBeta(phi = phi, delta_t = 1:5)
#>
#> Total, Direct, and Indirect Effects
#>
#> effect interval est se R 2.5% 97.5%
#> 1 from x to x 1 0.6998 0.0463 1000 0.6151 0.7991
#> 2 from x to m 1 0.4977 0.0345 1000 0.4344 0.5679
#> 3 from x to y 1 -0.1005 0.0307 1000 -0.1602 -0.0410
#> 4 from m to x 1 0.0021 0.0439 1000 -0.0829 0.0894
#> 5 from m to m 1 0.6013 0.0322 1000 0.5393 0.6640
#> 6 from m to y 1 0.3994 0.0283 1000 0.3447 0.4551
#> 7 from y to x 1 0.0009 0.0427 1000 -0.0843 0.0824
#> 8 from y to m 1 0.0000 0.0307 1000 -0.0589 0.0598
#> 9 from y to y 1 0.4997 0.0265 1000 0.4500 0.5523
#> 10 from x to x 2 0.4907 0.0543 1000 0.3994 0.6117
#> 11 from x to m 2 0.6476 0.0525 1000 0.5522 0.7579
#> 12 from x to y 2 0.0783 0.0347 1000 0.0050 0.1411
#> 13 from m to x 2 0.0031 0.0515 1000 -0.0970 0.1046
#> 14 from m to m 2 0.3626 0.0493 1000 0.2690 0.4606
#> 15 from m to y 2 0.4395 0.0332 1000 0.3789 0.5078
#> 16 from y to x 2 0.0011 0.0516 1000 -0.1025 0.0999
#> 17 from y to m 2 0.0004 0.0497 1000 -0.1008 0.0965
#> 18 from y to y 2 0.2496 0.0306 1000 0.1939 0.3104
#> 19 from x to x 3 0.3448 0.0547 1000 0.2528 0.4631
#> 20 from x to m 3 0.6336 0.0647 1000 0.5207 0.7680
#> 21 from x to y 3 0.2484 0.0356 1000 0.1777 0.3183
#> 22 from m to x 3 0.0034 0.0500 1000 -0.0930 0.1029
#> 23 from m to m 3 0.2195 0.0594 1000 0.1071 0.3407
#> 24 from m to y 3 0.3641 0.0335 1000 0.3028 0.4321
#> 25 from y to x 3 0.0010 0.0473 1000 -0.0955 0.0913
#> 26 from y to m 3 0.0008 0.0598 1000 -0.1181 0.1113
#> 27 from y to y 3 0.1248 0.0288 1000 0.0711 0.1838
#> 28 from x to x 4 0.2428 0.0539 1000 0.1526 0.3541
#> 29 from x to m 4 0.5526 0.0716 1000 0.4337 0.7058
#> 30 from x to y 4 0.3425 0.0398 1000 0.2701 0.4224
#> 31 from m to x 4 0.0031 0.0461 1000 -0.0876 0.0980
#> 32 from m to m 4 0.1337 0.0636 1000 0.0143 0.2657
#> 33 from m to y 4 0.2693 0.0351 1000 0.2060 0.3392
#> 34 from y to x 4 0.0008 0.0391 1000 -0.0804 0.0731
#> 35 from y to m 4 0.0009 0.0617 1000 -0.1224 0.1118
#> 36 from y to y 4 0.0626 0.0304 1000 0.0032 0.1251
#> 37 from x to x 5 0.1714 0.0531 1000 0.0838 0.2841
#> 38 from x to m 5 0.4531 0.0749 1000 0.3274 0.6108
#> 39 from x to y 5 0.3674 0.0449 1000 0.2855 0.4639
#> 40 from m to x 5 0.0027 0.0411 1000 -0.0776 0.0890
#> 41 from m to m 5 0.0819 0.0635 1000 -0.0414 0.2158
#> 42 from m to y 5 0.1876 0.0374 1000 0.1198 0.2662
#> 43 from y to x 5 0.0006 0.0307 1000 -0.0638 0.0584
#> 44 from y to m 5 0.0010 0.0577 1000 -0.1192 0.1073
#> 45 from y to y 5 0.0316 0.0342 1000 -0.0349 0.0988
confint(posterior, level = 0.95)
#> effect interval 2.5 % 97.5 %
#> 1 from x to x 1 0.615077559 0.79909237
#> 2 from x to m 1 0.434406622 0.56791660
#> 3 from x to y 1 -0.160219435 -0.04098514
#> 4 from x to x 2 0.399419871 0.61173690
#> 5 from x to m 2 0.552153140 0.75786017
#> 6 from x to y 2 0.004972431 0.14110172
#> 7 from x to x 3 0.252788824 0.46310472
#> 8 from x to m 3 0.520666214 0.76798977
#> 9 from x to y 3 0.177739549 0.31828824
#> 10 from x to x 4 0.152597586 0.35410705
#> 11 from x to m 4 0.433719879 0.70584106
#> 12 from x to y 4 0.270116254 0.42244066
#> 13 from x to x 5 0.083759930 0.28405495
#> 14 from x to m 5 0.327425315 0.61079299
#> 15 from x to y 5 0.285480283 0.46391752
plot(posterior)
# Methods -------------------------------------------------------------------
# PosteriorBeta has a number of methods including
# print, summary, confint, and plot
print(posterior)
#> Call:
#> PosteriorBeta(phi = phi, delta_t = 1:5)
#>
#> Total, Direct, and Indirect Effects
#>
#> effect interval est se R 2.5% 97.5%
#> 1 from x to x 1 0.6998 0.0463 1000 0.6151 0.7991
#> 2 from x to m 1 0.4977 0.0345 1000 0.4344 0.5679
#> 3 from x to y 1 -0.1005 0.0307 1000 -0.1602 -0.0410
#> 4 from m to x 1 0.0021 0.0439 1000 -0.0829 0.0894
#> 5 from m to m 1 0.6013 0.0322 1000 0.5393 0.6640
#> 6 from m to y 1 0.3994 0.0283 1000 0.3447 0.4551
#> 7 from y to x 1 0.0009 0.0427 1000 -0.0843 0.0824
#> 8 from y to m 1 0.0000 0.0307 1000 -0.0589 0.0598
#> 9 from y to y 1 0.4997 0.0265 1000 0.4500 0.5523
#> 10 from x to x 2 0.4907 0.0543 1000 0.3994 0.6117
#> 11 from x to m 2 0.6476 0.0525 1000 0.5522 0.7579
#> 12 from x to y 2 0.0783 0.0347 1000 0.0050 0.1411
#> 13 from m to x 2 0.0031 0.0515 1000 -0.0970 0.1046
#> 14 from m to m 2 0.3626 0.0493 1000 0.2690 0.4606
#> 15 from m to y 2 0.4395 0.0332 1000 0.3789 0.5078
#> 16 from y to x 2 0.0011 0.0516 1000 -0.1025 0.0999
#> 17 from y to m 2 0.0004 0.0497 1000 -0.1008 0.0965
#> 18 from y to y 2 0.2496 0.0306 1000 0.1939 0.3104
#> 19 from x to x 3 0.3448 0.0547 1000 0.2528 0.4631
#> 20 from x to m 3 0.6336 0.0647 1000 0.5207 0.7680
#> 21 from x to y 3 0.2484 0.0356 1000 0.1777 0.3183
#> 22 from m to x 3 0.0034 0.0500 1000 -0.0930 0.1029
#> 23 from m to m 3 0.2195 0.0594 1000 0.1071 0.3407
#> 24 from m to y 3 0.3641 0.0335 1000 0.3028 0.4321
#> 25 from y to x 3 0.0010 0.0473 1000 -0.0955 0.0913
#> 26 from y to m 3 0.0008 0.0598 1000 -0.1181 0.1113
#> 27 from y to y 3 0.1248 0.0288 1000 0.0711 0.1838
#> 28 from x to x 4 0.2428 0.0539 1000 0.1526 0.3541
#> 29 from x to m 4 0.5526 0.0716 1000 0.4337 0.7058
#> 30 from x to y 4 0.3425 0.0398 1000 0.2701 0.4224
#> 31 from m to x 4 0.0031 0.0461 1000 -0.0876 0.0980
#> 32 from m to m 4 0.1337 0.0636 1000 0.0143 0.2657
#> 33 from m to y 4 0.2693 0.0351 1000 0.2060 0.3392
#> 34 from y to x 4 0.0008 0.0391 1000 -0.0804 0.0731
#> 35 from y to m 4 0.0009 0.0617 1000 -0.1224 0.1118
#> 36 from y to y 4 0.0626 0.0304 1000 0.0032 0.1251
#> 37 from x to x 5 0.1714 0.0531 1000 0.0838 0.2841
#> 38 from x to m 5 0.4531 0.0749 1000 0.3274 0.6108
#> 39 from x to y 5 0.3674 0.0449 1000 0.2855 0.4639
#> 40 from m to x 5 0.0027 0.0411 1000 -0.0776 0.0890
#> 41 from m to m 5 0.0819 0.0635 1000 -0.0414 0.2158
#> 42 from m to y 5 0.1876 0.0374 1000 0.1198 0.2662
#> 43 from y to x 5 0.0006 0.0307 1000 -0.0638 0.0584
#> 44 from y to m 5 0.0010 0.0577 1000 -0.1192 0.1073
#> 45 from y to y 5 0.0316 0.0342 1000 -0.0349 0.0988
summary(posterior)
#> Call:
#> PosteriorBeta(phi = phi, delta_t = 1:5)
#>
#> Total, Direct, and Indirect Effects
#>
#> effect interval est se R 2.5% 97.5%
#> 1 from x to x 1 0.6998 0.0463 1000 0.6151 0.7991
#> 2 from x to m 1 0.4977 0.0345 1000 0.4344 0.5679
#> 3 from x to y 1 -0.1005 0.0307 1000 -0.1602 -0.0410
#> 4 from m to x 1 0.0021 0.0439 1000 -0.0829 0.0894
#> 5 from m to m 1 0.6013 0.0322 1000 0.5393 0.6640
#> 6 from m to y 1 0.3994 0.0283 1000 0.3447 0.4551
#> 7 from y to x 1 0.0009 0.0427 1000 -0.0843 0.0824
#> 8 from y to m 1 0.0000 0.0307 1000 -0.0589 0.0598
#> 9 from y to y 1 0.4997 0.0265 1000 0.4500 0.5523
#> 10 from x to x 2 0.4907 0.0543 1000 0.3994 0.6117
#> 11 from x to m 2 0.6476 0.0525 1000 0.5522 0.7579
#> 12 from x to y 2 0.0783 0.0347 1000 0.0050 0.1411
#> 13 from m to x 2 0.0031 0.0515 1000 -0.0970 0.1046
#> 14 from m to m 2 0.3626 0.0493 1000 0.2690 0.4606
#> 15 from m to y 2 0.4395 0.0332 1000 0.3789 0.5078
#> 16 from y to x 2 0.0011 0.0516 1000 -0.1025 0.0999
#> 17 from y to m 2 0.0004 0.0497 1000 -0.1008 0.0965
#> 18 from y to y 2 0.2496 0.0306 1000 0.1939 0.3104
#> 19 from x to x 3 0.3448 0.0547 1000 0.2528 0.4631
#> 20 from x to m 3 0.6336 0.0647 1000 0.5207 0.7680
#> 21 from x to y 3 0.2484 0.0356 1000 0.1777 0.3183
#> 22 from m to x 3 0.0034 0.0500 1000 -0.0930 0.1029
#> 23 from m to m 3 0.2195 0.0594 1000 0.1071 0.3407
#> 24 from m to y 3 0.3641 0.0335 1000 0.3028 0.4321
#> 25 from y to x 3 0.0010 0.0473 1000 -0.0955 0.0913
#> 26 from y to m 3 0.0008 0.0598 1000 -0.1181 0.1113
#> 27 from y to y 3 0.1248 0.0288 1000 0.0711 0.1838
#> 28 from x to x 4 0.2428 0.0539 1000 0.1526 0.3541
#> 29 from x to m 4 0.5526 0.0716 1000 0.4337 0.7058
#> 30 from x to y 4 0.3425 0.0398 1000 0.2701 0.4224
#> 31 from m to x 4 0.0031 0.0461 1000 -0.0876 0.0980
#> 32 from m to m 4 0.1337 0.0636 1000 0.0143 0.2657
#> 33 from m to y 4 0.2693 0.0351 1000 0.2060 0.3392
#> 34 from y to x 4 0.0008 0.0391 1000 -0.0804 0.0731
#> 35 from y to m 4 0.0009 0.0617 1000 -0.1224 0.1118
#> 36 from y to y 4 0.0626 0.0304 1000 0.0032 0.1251
#> 37 from x to x 5 0.1714 0.0531 1000 0.0838 0.2841
#> 38 from x to m 5 0.4531 0.0749 1000 0.3274 0.6108
#> 39 from x to y 5 0.3674 0.0449 1000 0.2855 0.4639
#> 40 from m to x 5 0.0027 0.0411 1000 -0.0776 0.0890
#> 41 from m to m 5 0.0819 0.0635 1000 -0.0414 0.2158
#> 42 from m to y 5 0.1876 0.0374 1000 0.1198 0.2662
#> 43 from y to x 5 0.0006 0.0307 1000 -0.0638 0.0584
#> 44 from y to m 5 0.0010 0.0577 1000 -0.1192 0.1073
#> 45 from y to y 5 0.0316 0.0342 1000 -0.0349 0.0988
confint(posterior, level = 0.95)
#> effect interval 2.5 % 97.5 %
#> 1 from x to x 1 0.615077559 0.79909237
#> 2 from x to m 1 0.434406622 0.56791660
#> 3 from x to y 1 -0.160219435 -0.04098514
#> 4 from x to x 2 0.399419871 0.61173690
#> 5 from x to m 2 0.552153140 0.75786017
#> 6 from x to y 2 0.004972431 0.14110172
#> 7 from x to x 3 0.252788824 0.46310472
#> 8 from x to m 3 0.520666214 0.76798977
#> 9 from x to y 3 0.177739549 0.31828824
#> 10 from x to x 4 0.152597586 0.35410705
#> 11 from x to m 4 0.433719879 0.70584106
#> 12 from x to y 4 0.270116254 0.42244066
#> 13 from x to x 5 0.083759930 0.28405495
#> 14 from x to m 5 0.327425315 0.61079299
#> 15 from x to y 5 0.285480283 0.46391752
plot(posterior)