Benchmark: Comparing the Monte Carlo Method with Nonparametric Bootstrapping (MI)
Ivan Jacob Agaloos Pesigan
2026-03-01
Source:vignettes/benchmark-mi.Rmd
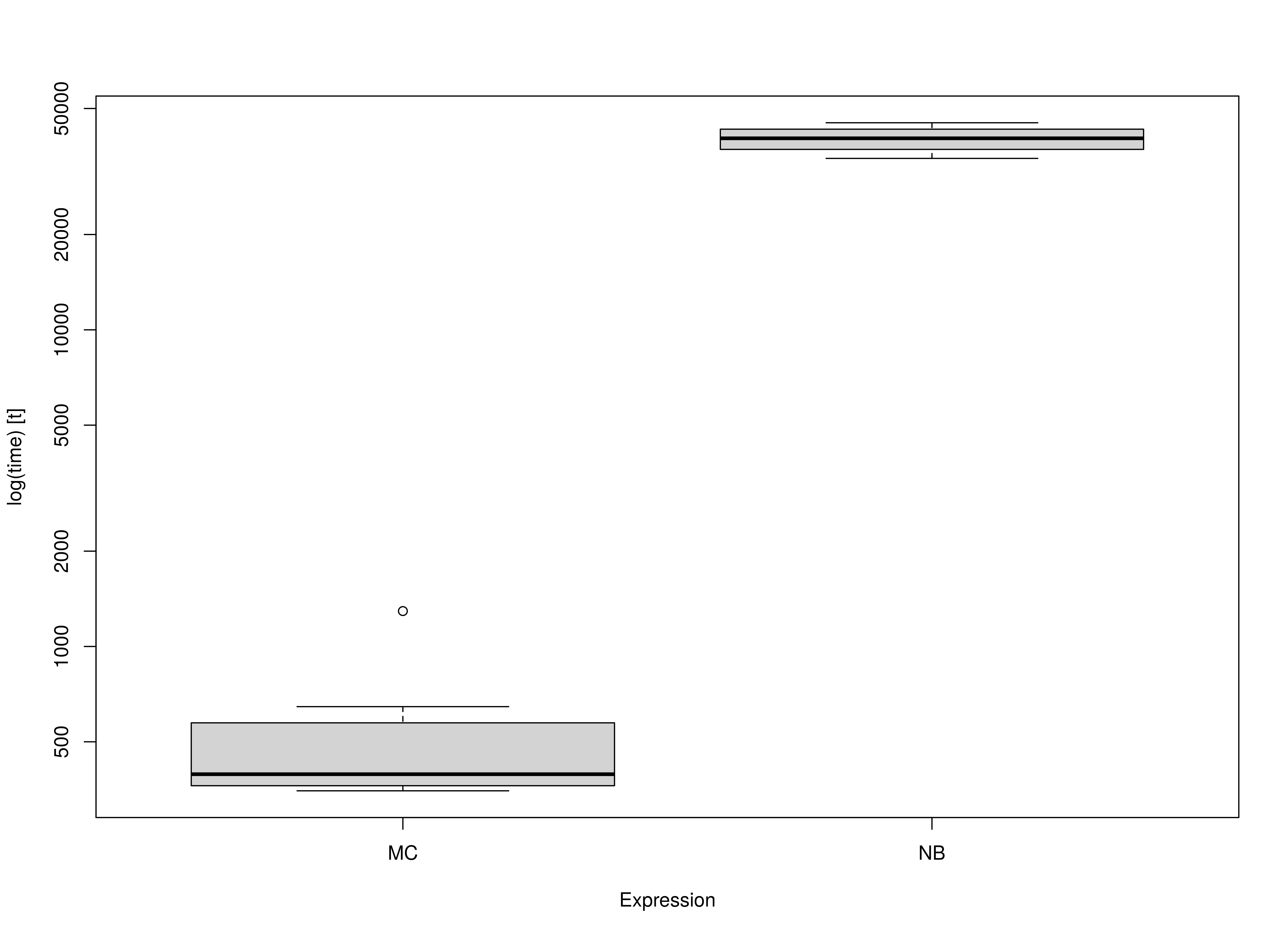
benchmark-mi.RmdWe compare the Monte Carlo (MC) method with nonparametric bootstrapping (NB) using the simple mediation model with missing data using multiple imputation. One advantage of MC over NB is speed. This is because the model is only fitted once in MC whereas it is fitted many times in NB.
Data
n <- 1000
a <- 0.50
b <- 0.50
cp <- 0.25
s2_em <- 1 - a^2
s2_ey <- 1 - cp^2 - a^2 * b^2 - b^2 * s2_em - 2 * cp * a * b
em <- rnorm(n = n, mean = 0, sd = sqrt(s2_em))
ey <- rnorm(n = n, mean = 0, sd = sqrt(s2_ey))
X <- rnorm(n = n)
M <- a * X + em
Y <- cp * X + b * M + ey
df <- data.frame(X, M, Y)
# Create data set with missing values.
miss <- sample(1:dim(df)[1], 300)
df[miss[1:100], "X"] <- NA
df[miss[101:200], "M"] <- NA
df[miss[201:300], "Y"] <- NAMultiple Imputation
Perform the appropriate multiple imputation approach to deal with missing values. In this example, we impute multivariate missing data under the normal model.
mi <- amelia(
x = df,
m = 5L,
p2s = 0
)Model Specification
The indirect effect is defined by the product of the slopes of paths
X to M labeled as a and
M to Y labeled as b. In this
example, we are interested in the confidence intervals of
indirect defined as the product of a and
b using the := operator in the
lavaan model syntax.
model <- "
Y ~ cp * X + b * M
M ~ a * X
X ~~ X
indirect := a * b
direct := cp
total := cp + (a * b)
"Model Fitting
We can now fit the model using the sem() function from
lavaan. We do not need to deal with missing values in this
stage.
fit <- sem(data = df, model = model)Monte Carlo Confidence Intervals (Multiple Imputation)
The fit lavaan object and mi
mids object can then be passed to the MCMI()
function from semmcci to generate Monte Carlo confidence
intervals using multiple imputation as described in Pesigan and Cheung
(2024).
MCMI(fit, R = 100L, alpha = 0.05, mi = mi)
#> Monte Carlo Confidence Intervals (Multiple Imputation Estimates)
#> est se R 2.5% 97.5%
#> cp 0.2274 0.0295 100 0.1787 0.2818
#> b 0.5192 0.0342 100 0.4534 0.5839
#> a 0.4790 0.0281 100 0.4249 0.5266
#> X~~X 1.0613 0.0443 100 0.9775 1.1328
#> Y~~Y 0.5439 0.0244 100 0.5010 0.5911
#> M~~M 0.7642 0.0397 100 0.7048 0.8397
#> indirect 0.2486 0.0189 100 0.2103 0.2755
#> direct 0.2274 0.0295 100 0.1787 0.2818
#> total 0.4760 0.0288 100 0.4243 0.5329Nonparametric Bootstrap Confidence Intervals (Multiple Imputation)
Nonparametric bootstrap confidence intervals can be generated in
bmemLavaan using the following.
summary(
bmemLavaan::bmem(data = df, model = model, method = "mi", boot = 100L, m = 5L)
)
#>
#> Estimate method: multiple imputation
#> Sample size: 1000
#> Number of request bootstrap draws: 100
#> Number of successful bootstrap draws: 100
#> Type of confidence interval: perc
#>
#> Values of statistics:
#>
#> Value SE 2.5% 97.5%
#> chisq 0.000 0.000 0.000 0.000
#> GFI 1.000 0.000 1.000 1.000
#> AGFI 1.000 0.000 1.000 1.000
#> RMSEA 0.000 0.000 0.000 0.000
#> NFI 1.000 0.000 1.000 1.000
#> NNFI 1.000 0.000 1.000 1.000
#> CFI 1.000 0.000 1.000 1.000
#> BIC 7742.967 81.777 7575.258 7857.675
#> SRMR 0.000 0.000 0.000 0.000
#>
#> Estimation of parameters:
#>
#> Estimate SE 2.5% 97.5%
#> Regressions:
#> Y ~
#> X (cp) 0.234 0.030 0.176 0.296
#> M (b) 0.513 0.032 0.460 0.570
#> M ~
#> X (a) 0.476 0.030 0.426 0.540
#>
#> Variances:
#> X 1.057 0.046 0.950 1.144
#> Y 0.556 0.027 0.488 0.600
#> M 0.755 0.035 0.684 0.813
#>
#>
#>
#> Defined parameters:
#> a*b (indr) 0.244 0.020 0.206 0.285
#> cp (drct) 0.234 0.030 0.176 0.296
#> cp+(*) (totl) 0.479 0.030 0.428 0.539Benchmark
benchmark_mi_01 <- microbenchmark(
MC = {
fit <- sem(
data = df,
model = model
)
mi <- Amelia::amelia(
x = df,
m = m,
p2s = 0
)
MCMI(
fit,
R = R,
decomposition = "chol",
pd = FALSE,
mi = mi
)
},
NB = bmemLavaan::bmem(
data = df,
model = model,
method = "mi",
boot = B,
m = m
),
times = 10
)Summary of Benchmark Results
summary(benchmark_mi_01, unit = "ms")
#> expr min lq mean median uq max neval
#> 1 MC 714.9348 719.4493 753.607 737.44 746.9044 933.4056 10
#> 2 NB 71749.4791 72082.4629 72354.947 72310.45 72488.2285 73271.0202 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_mi_01, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.0000 1.0000 1.00000 1.00000 1.00000 1.00000 10
#> 2 NB 100.3581 100.1912 96.01151 98.05604 97.05155 78.49859 10Benchmark - Monte Carlo Method with Precalculated Estimates and Multiple Imputation
fit <- sem(
data = df,
model = model
)
mi <- Amelia::amelia(
x = df,
m = m,
p2s = 0
)
benchmark_mi_02 <- microbenchmark(
MC = MCMI(
fit,
R = R,
decomposition = "chol",
pd = FALSE,
mi = mi
),
NB = bmemLavaan::bmem(
data = df,
model = model,
method = "mi",
boot = B,
m = m
),
times = 10
)Summary of Benchmark Results
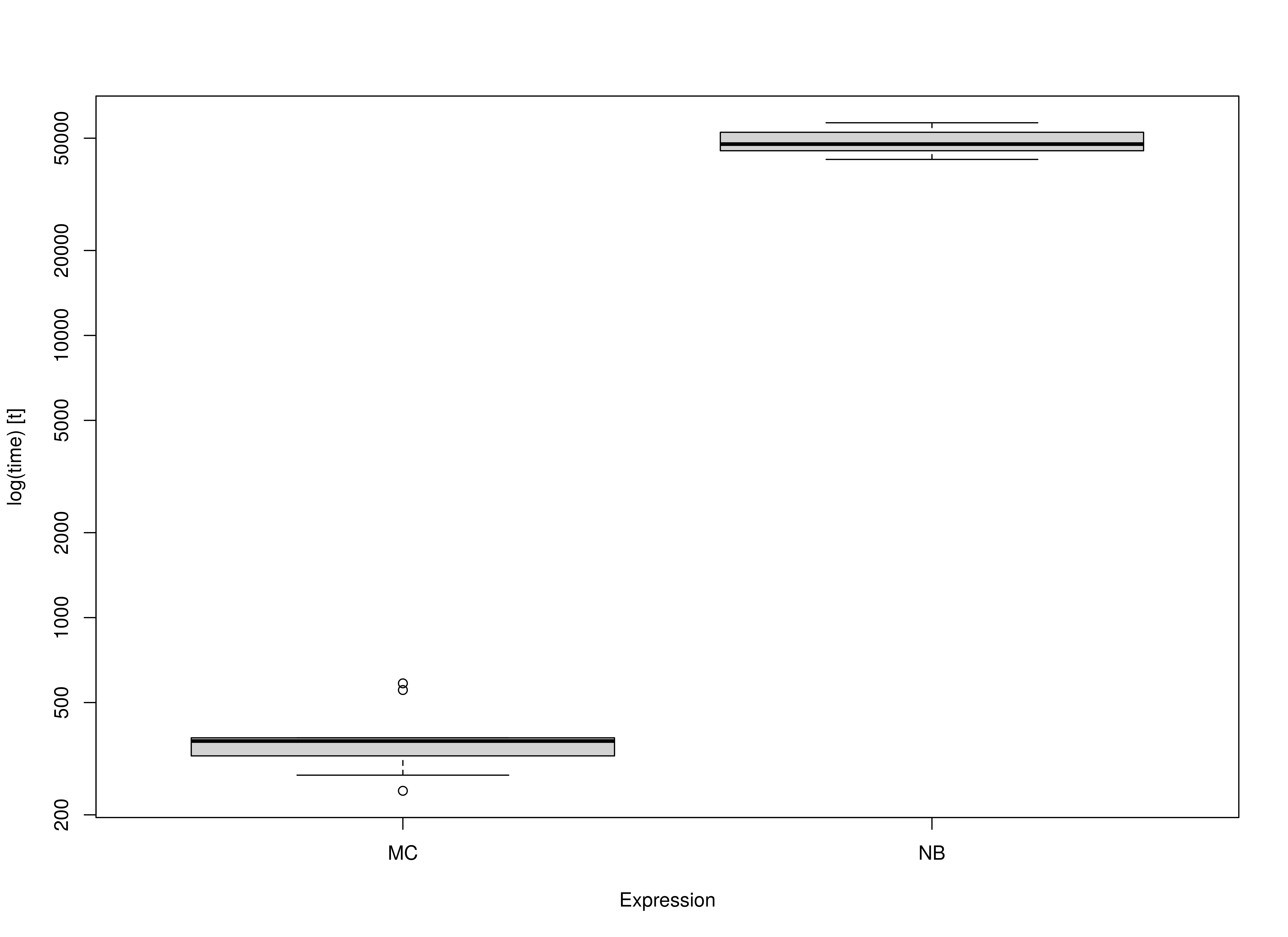
summary(benchmark_mi_02, unit = "ms")
#> expr min lq mean median uq max neval
#> 1 MC 540.1188 543.1085 548.4491 547.7864 553.0569 558.0039 10
#> 2 NB 72096.5023 72292.0301 72413.8396 72434.4786 72575.3305 72588.6181 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_mi_02, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 10
#> 2 NB 133.4827 133.1079 132.0338 132.2313 131.2258 130.0862 10