Benchmark: Comparing the Monte Carlo Method with Nonparametric Bootstrapping (FIML)
Ivan Jacob Agaloos Pesigan
2026-03-01
Source:vignettes/benchmark-fiml.Rmd
benchmark-fiml.RmdWe compare the Monte Carlo (MC) method with nonparametric bootstrapping (NB) using the simple mediation model with missing data using full-information maximum likelihood. One advantage of MC over NB is speed. This is because the model is only fitted once in MC whereas it is fitted many times in NB.
Data
n <- 1000
a <- 0.50
b <- 0.50
cp <- 0.25
s2_em <- 1 - a^2
s2_ey <- 1 - cp^2 - a^2 * b^2 - b^2 * s2_em - 2 * cp * a * b
em <- rnorm(n = n, mean = 0, sd = sqrt(s2_em))
ey <- rnorm(n = n, mean = 0, sd = sqrt(s2_ey))
X <- rnorm(n = n)
M <- a * X + em
Y <- cp * X + b * M + ey
df <- data.frame(X, M, Y)
# Create data set with missing values.
miss <- sample(1:dim(df)[1], 300)
df[miss[1:100], "X"] <- NA
df[miss[101:200], "M"] <- NA
df[miss[201:300], "Y"] <- NAModel Specification
The indirect effect is defined by the product of the slopes of paths
X to M labeled as a and
M to Y labeled as b. In this
example, we are interested in the confidence intervals of
indirect defined as the product of a and
b using the := operator in the
lavaan model syntax.
model <- "
Y ~ cp * X + b * M
M ~ a * X
X ~~ X
indirect := a * b
direct := cp
total := cp + (a * b)
"Model Fitting
We can now fit the model using the sem() function from
lavaan. We are using missing = "fiml" to
handle missing data in lavaan.
fit <- sem(data = df, model = model)Monte Carlo Confidence Intervals
The fit lavaan object can then be passed to
the MC() function from semmcci to generate
Monte Carlo confidence intervals.
MC(fit, R = 100L, alpha = 0.05)
#> Monte Carlo Confidence Intervals
#> est se R 2.5% 97.5%
#> cp 0.2419 0.0332 100 0.1792 0.3070
#> b 0.5166 0.0308 100 0.4580 0.5785
#> a 0.4989 0.0319 100 0.4448 0.5615
#> X~~X 1.0951 0.0621 100 0.9875 1.2045
#> Y~~Y 0.5796 0.0307 100 0.5179 0.6336
#> M~~M 0.8045 0.0464 100 0.7325 0.9106
#> indirect 0.2577 0.0210 100 0.2234 0.3031
#> direct 0.2419 0.0332 100 0.1792 0.3070
#> total 0.4996 0.0322 100 0.4550 0.5681Nonparametric Bootstrap Confidence Intervals
Nonparametric bootstrap confidence intervals can be generated in
lavaan using the following.
parameterEstimates(
sem(
data = df,
model = model,
missing = "fiml",
se = "bootstrap",
bootstrap = 100L
)
)
#> lhs op rhs label est se z pvalue ci.lower ci.upper
#> 1 Y ~ X cp 0.234 0.030 7.721 0.000 0.169 0.287
#> 2 Y ~ M b 0.511 0.035 14.704 0.000 0.442 0.585
#> 3 M ~ X a 0.481 0.028 17.117 0.000 0.425 0.532
#> 4 X ~~ X 1.059 0.049 21.539 0.000 0.979 1.148
#> 5 Y ~~ Y 0.554 0.029 19.264 0.000 0.490 0.607
#> 6 M ~~ M 0.756 0.032 23.389 0.000 0.693 0.820
#> 7 Y ~1 -0.013 0.027 -0.473 0.636 -0.065 0.056
#> 8 M ~1 -0.022 0.030 -0.744 0.457 -0.077 0.044
#> 9 X ~1 0.002 0.036 0.069 0.945 -0.072 0.074
#> 10 indirect := a*b indirect 0.246 0.021 11.534 0.000 0.202 0.286
#> 11 direct := cp direct 0.234 0.030 7.721 0.000 0.169 0.287
#> 12 total := cp+(a*b) total 0.479 0.030 16.081 0.000 0.417 0.547Benchmark
Arguments
| Variables | Values | Notes |
|---|---|---|
| R | 1000 | Number of Monte Carlo replications. |
| B | 1000 | Number of bootstrap samples. |
benchmark_fiml_01 <- microbenchmark(
MC = {
fit <- sem(
data = df,
model = model,
missing = "fiml"
)
MC(
fit,
R = R,
decomposition = "chol",
pd = FALSE
)
},
NB = sem(
data = df,
model = model,
missing = "fiml",
se = "bootstrap",
bootstrap = B
),
times = 10
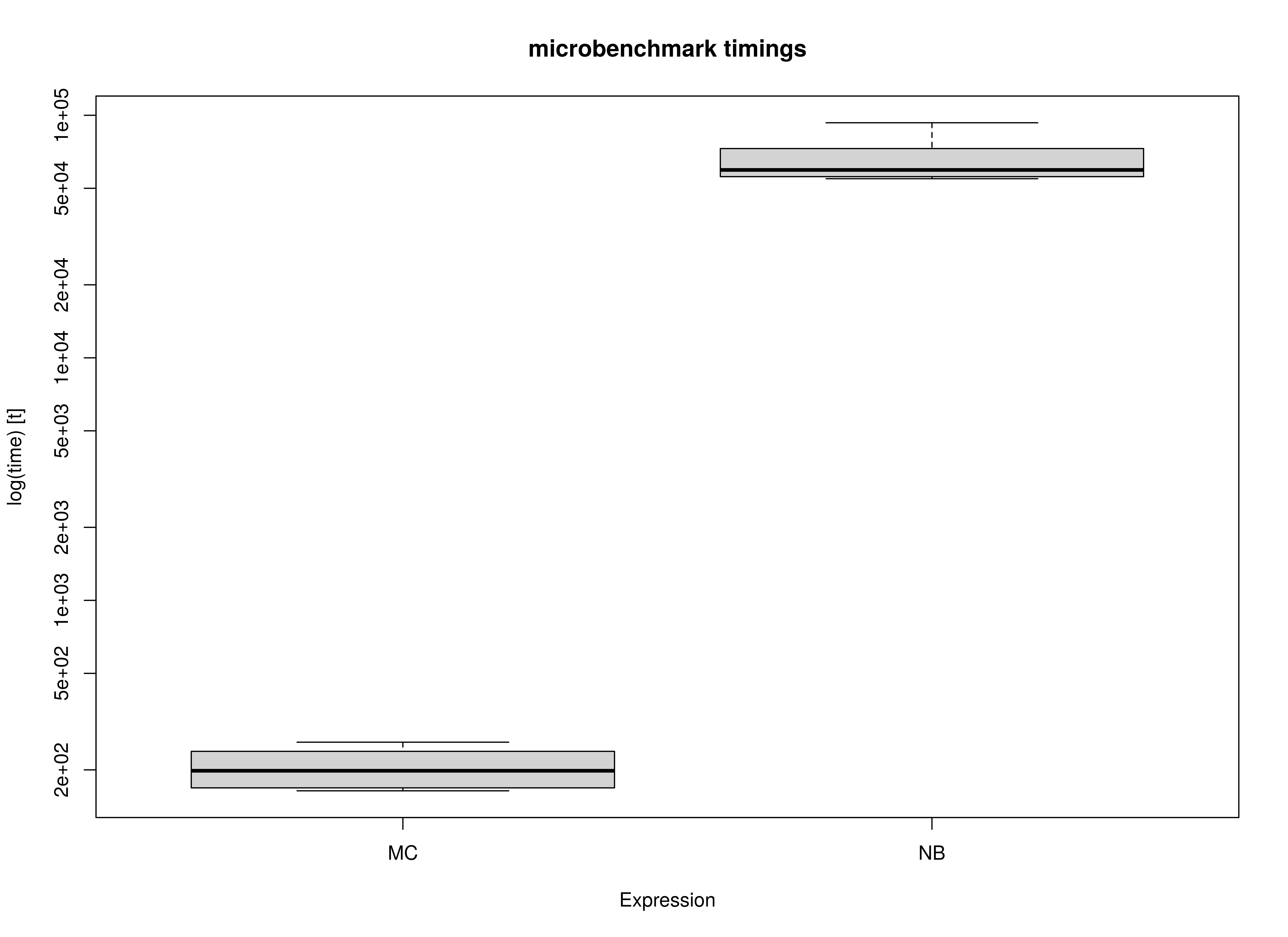
)Summary of Benchmark Results
summary(benchmark_fiml_01, unit = "ms")
#> expr min lq mean median uq max neval
#> 1 MC 260.1772 260.9986 261.2316 261.2154 261.5805 261.8205 10
#> 2 NB 60196.0501 60273.7339 60304.4039 60293.9547 60347.5345 60434.7181 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_fiml_01, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.0000 1.0000 1.0000 1.0000 1.0000 1.000 10
#> 2 NB 231.3656 230.9351 230.8465 230.8208 230.7035 230.825 10Benchmark - Monte Carlo Method with Precalculated Estimates
fit <- sem(
data = df,
model = model,
missing = "fiml"
)
benchmark_fiml_02 <- microbenchmark(
MC = MC(
fit,
R = R,
decomposition = "chol",
pd = FALSE
),
NB = sem(
data = df,
model = model,
missing = "fiml",
se = "bootstrap",
bootstrap = B
),
times = 10
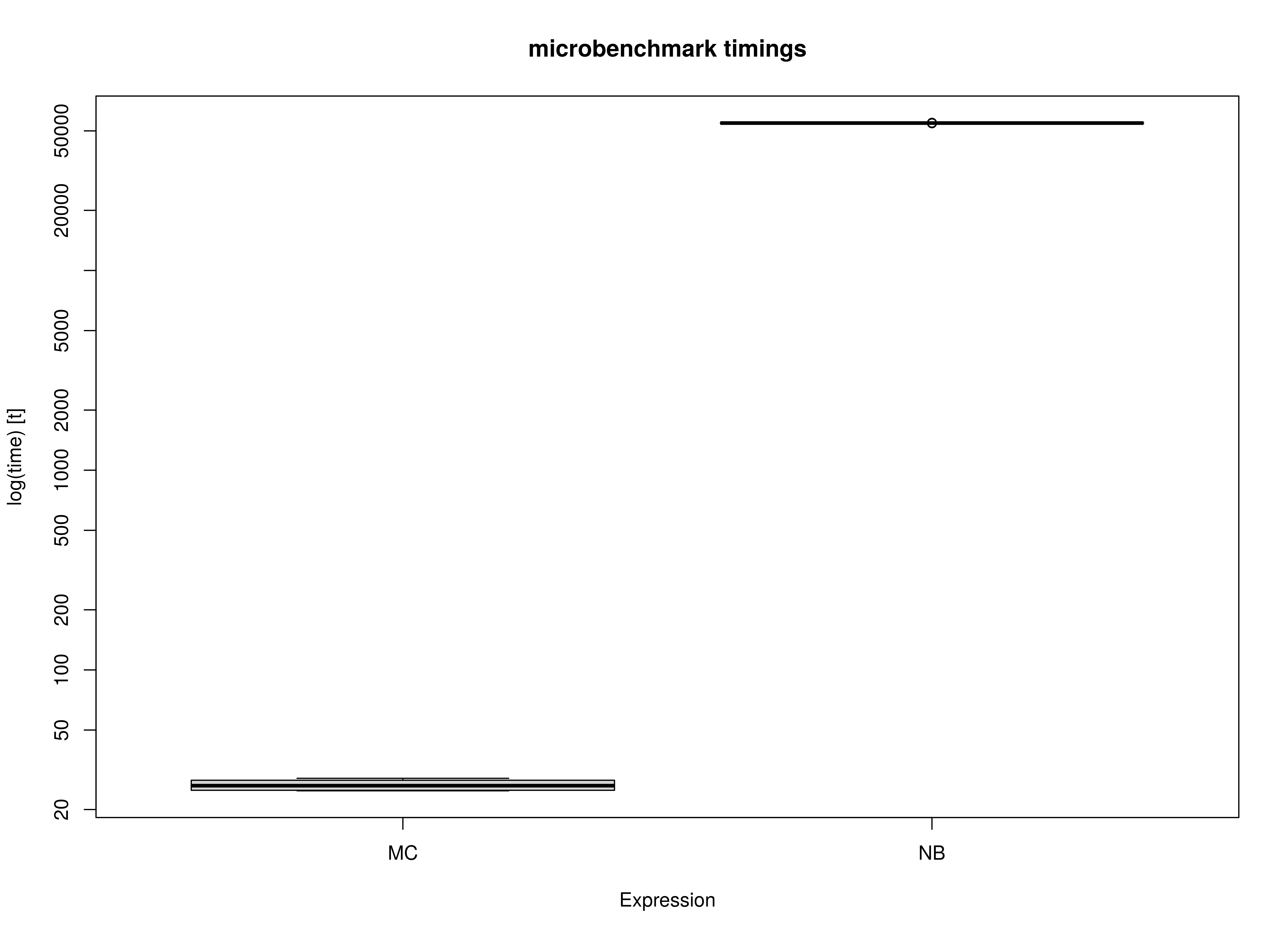
)Summary of Benchmark Results
summary(benchmark_fiml_02, unit = "ms")
#> expr min lq mean median uq max
#> 1 MC 58.32543 58.44053 60.16606 59.65633 60.9178 63.67535
#> 2 NB 60136.15726 60200.18985 60245.00599 60241.29127 60281.3703 60384.01692
#> neval
#> 1 10
#> 2 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_fiml_02, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.000 1.00 1.000 1.000 1.0000 1.0000 10
#> 2 NB 1031.045 1030.11 1001.312 1009.806 989.5526 948.3108 10