Benchmark: Comparing the Monte Carlo Method with Nonparametric Bootstrapping
Ivan Jacob Agaloos Pesigan
2026-03-01
Source:vignettes/benchmark-complete.Rmd
benchmark-complete.RmdWe compare the Monte Carlo (MC) method with nonparametric bootstrapping (NB) using the simple mediation model with complete data. One advantage of MC over NB is speed. This is because the model is only fitted once in MC whereas it is fitted many times in NB.
Model Specification
The indirect effect is defined by the product of the slopes of paths
X to M labeled as a and
M to Y labeled as b. In this
example, we are interested in the confidence intervals of
indirect defined as the product of a and
b using the := operator in the
lavaan model syntax.
model <- "
Y ~ cp * X + b * M
M ~ a * X
X ~~ X
indirect := a * b
direct := cp
total := cp + (a * b)
"Model Fitting
We can now fit the model using the sem() function from
lavaan.
fit <- sem(data = df, model = model)Monte Carlo Confidence Intervals
The fit lavaan object can then be passed to
the MC() function from semmcci to generate
Monte Carlo confidence intervals.
MC(fit, R = 100L, alpha = 0.05)
#> Monte Carlo Confidence Intervals
#> est se R 2.5% 97.5%
#> cp 0.2333 0.0296 100 0.1806 0.2903
#> b 0.5082 0.0279 100 0.4555 0.5527
#> a 0.4820 0.0280 100 0.4220 0.5301
#> X~~X 1.0590 0.0426 100 0.9883 1.1428
#> Y~~Y 0.5462 0.0231 100 0.5064 0.5959
#> M~~M 0.7527 0.0337 100 0.6846 0.8029
#> indirect 0.2449 0.0179 100 0.2058 0.2738
#> direct 0.2333 0.0296 100 0.1806 0.2903
#> total 0.4782 0.0295 100 0.4162 0.5283Nonparametric Bootstrap Confidence Intervals
Nonparametric bootstrap confidence intervals can be generated in
lavaan using the following.
parameterEstimates(
sem(
data = df,
model = model,
se = "bootstrap",
bootstrap = 100L
)
)
#> lhs op rhs label est se z pvalue ci.lower ci.upper
#> 1 Y ~ X cp 0.233 0.025 9.395 0 0.183 0.278
#> 2 Y ~ M b 0.508 0.028 18.057 0 0.454 0.568
#> 3 M ~ X a 0.482 0.026 18.550 0 0.433 0.535
#> 4 X ~~ X 1.059 0.046 23.224 0 0.969 1.161
#> 5 Y ~~ Y 0.546 0.023 23.640 0 0.508 0.593
#> 6 M ~~ M 0.753 0.033 23.131 0 0.692 0.814
#> 7 indirect := a*b indirect 0.245 0.020 12.443 0 0.209 0.289
#> 8 direct := cp direct 0.233 0.025 9.395 0 0.183 0.278
#> 9 total := cp+(a*b) total 0.478 0.027 17.966 0 0.418 0.518Benchmark
Arguments
| Variables | Values | Notes |
|---|---|---|
| R | 1000 | Number of Monte Carlo replications. |
| B | 1000 | Number of bootstrap samples. |
benchmark_complete_01 <- microbenchmark(
MC = {
fit <- sem(
data = df,
model = model
)
MC(
fit,
R = R,
decomposition = "chol",
pd = FALSE
)
},
NB = sem(
data = df,
model = model,
se = "bootstrap",
bootstrap = B
),
times = 10
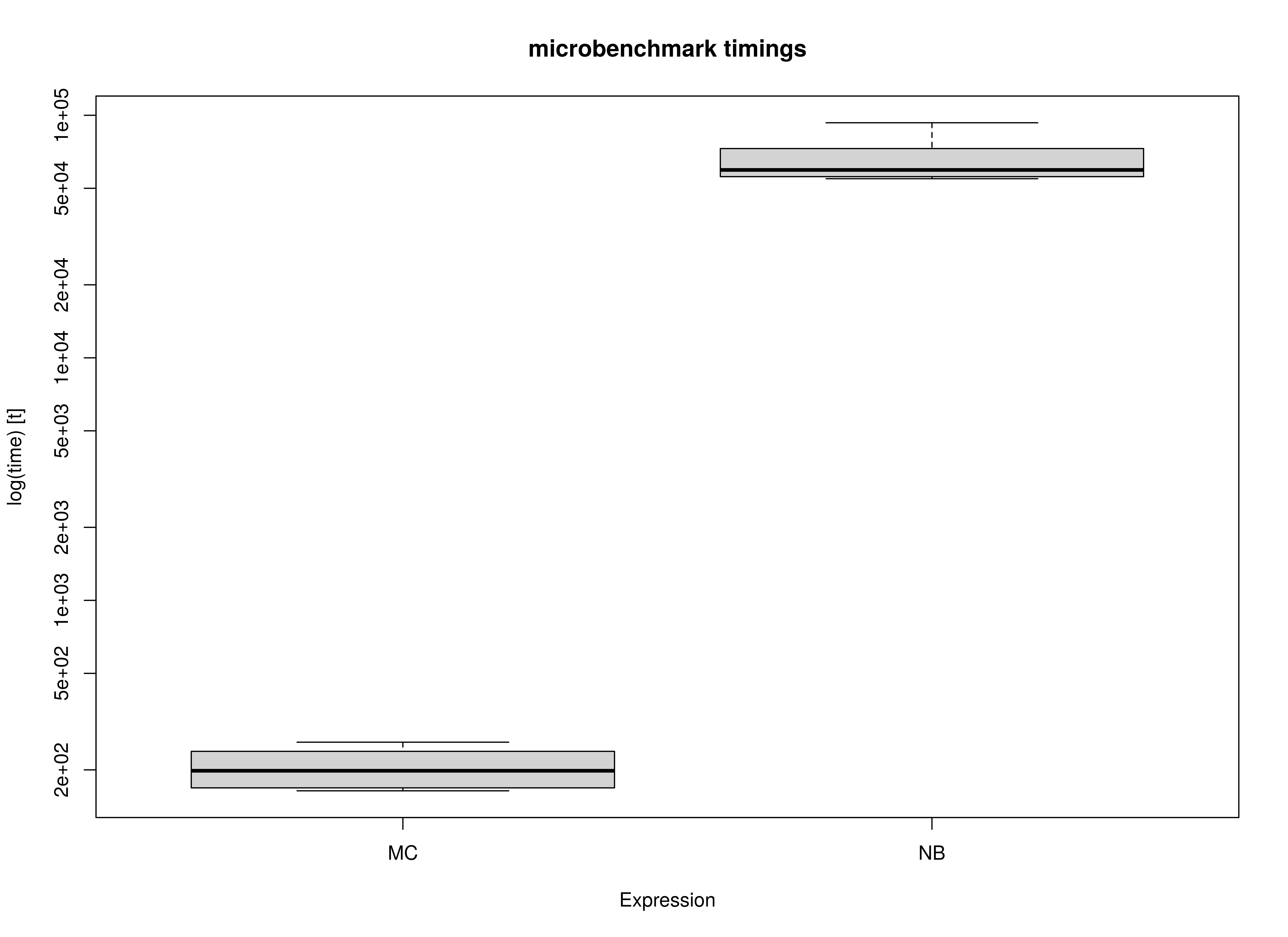
)Summary of Benchmark Results
summary(benchmark_complete_01, unit = "ms")
#> expr min lq mean median uq max neval
#> 1 MC 156.2475 163.342 178.0618 174.2547 177.4388 236.8905 10
#> 2 NB 28208.6459 29647.342 30674.5847 31136.5817 31477.8059 31873.9380 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_complete_01, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.0000 1.0000 1.0000 1.0000 1.0000 1.0000 10
#> 2 NB 180.5382 181.5047 172.2693 178.6843 177.4009 134.5514 10Benchmark - Monte Carlo Method with Precalculated Estimates
fit <- sem(
data = df,
model = model
)
benchmark_complete_02 <- microbenchmark(
MC = MC(
fit,
R = R,
decomposition = "chol",
pd = FALSE
),
NB = sem(
data = df,
model = model,
se = "bootstrap",
bootstrap = B
),
times = 10
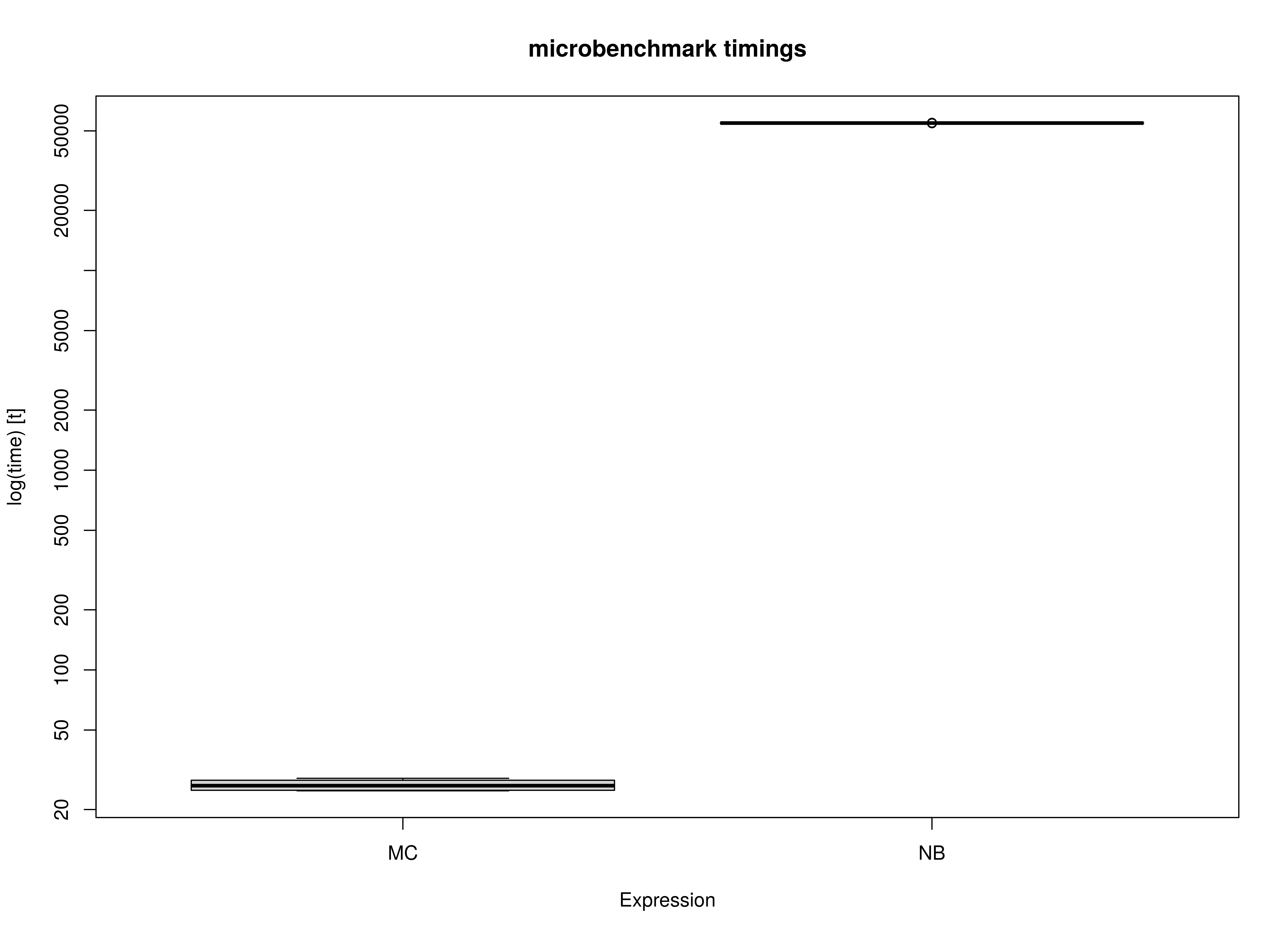
)Summary of Benchmark Results
summary(benchmark_complete_02, unit = "ms")
#> expr min lq mean median uq max
#> 1 MC 47.95931 50.42981 54.50973 55.05257 57.38705 62.35807
#> 2 NB 25714.54010 26012.19289 27636.27829 27725.97381 29009.61288 29622.84813
#> neval
#> 1 10
#> 2 10Summary of Benchmark Results Relative to the Faster Method
summary(benchmark_complete_02, unit = "relative")
#> expr min lq mean median uq max neval
#> 1 MC 1.0000 1.0000 1.0000 1.0000 1.000 1.0000 10
#> 2 NB 536.1741 515.8099 506.9971 503.6273 505.508 475.0443 10