Meta-Analysis of Discrete-Time Vector Autoregressive Model Estimates
Ivan Jacob Agaloos Pesigan
2026-05-08
Source:vignettes/meta-dt-var.Rmd
meta-dt-var.RmdDynamics Description
The Stable Reciprocal Regulation process represents a bivariate dynamic system in which two latent psychological constructs—such as negative and positive affect—mutually influence each other over time. Each construct shows moderate self-regulation (autoregressive effects) and mild opposing cross-effects, reflecting an equilibrium-seeking mechanism characteristic of emotional balance.
Individuals vary in their self-regulatory tendencies and in the strength of these antagonistic couplings. At the population level, the transition matrix indicates that increases in one construct are followed by slight decreases in the other, producing a stable, damped oscillatory pattern around individual equilibrium points. The process noise covariance allows for small correlated disturbances, while measurement errors are assumed to be minimal and symmetric across indicators.
This dynamic pattern captures a psychologically plausible process of reciprocal inhibition—where short-term fluctuations in one system component (e.g., negative affect) are naturally counteracted by adjustments in its counterpart (e.g., positive affect), leading to emotional homeostasis over time.
Model
The measurement model is given by where and are random variables.
The dynamic structure is given by where , , and are random variables, and , and are model parameters. Here, is a vector of latent variables at time and individual , represents a vector of latent variables at time and individual , and represents a vector of dynamic noise at time and individual . is a matrix of autoregression and cross regression coefficients for individual , and the covariance matrix of that is invariant across all individuals. In this model, is a symmetric matrix.
Data Generation
Notation
Let be the number of time points and be the number of individuals.
Let the initial condition be given by and are functions of and .
Let the set point vector be normally distributed with the following means and covariance matrix
Let the transition matrix be normally distributed with the following means and covariance matrix
The SimMuN and SimBetaN functions from the
simStateSpace package generate random set point vectors and
transition matrices from the multivariate normal distribution. Note that
the SimBetaN function generates transition matrices that
are weakly stationary with an option to set lower and upper bounds.
Let the dynamic process noise be given by
R Function Arguments
n
#> [1] 100
time
#> [1] 1000
# first mu0 in the list of length n
mu0[[1]]
#> [1] 2.287769 2.606098
# first sigma0 in the list of length n
sigma0[[1]]
#> [,1] [,2]
#> [1,] 1.4403549 0.5646226
#> [2,] 0.5646226 1.6885649
# first sigma0_l in the list of length n
sigma0_l[[1]] # sigma0_l <- t(chol(sigma0))
#> [,1] [,2]
#> [1,] 1.2001479 0.000000
#> [2,] 0.4704608 1.211293
# first alpha in the list of length n
alpha[[1]]
#> [1] 1.678968 2.010503
# first beta in the list of length n
beta[[1]]
#> [,1] [,2]
#> [1,] 0.32874206 -0.05498069
#> [2,] -0.07362662 0.29317209
# first psi in the list of length n
psi[[1]]
#> [,1] [,2]
#> [1,] 1.30 0.57
#> [2,] 0.57 1.56
psi_l[[1]] # psi_l <- t(chol(psi))
#> [,1] [,2]
#> [1,] 1.1401754 0.000000
#> [2,] 0.4999231 1.144586
# first mu (set point) in the list of length n
mu[[1]]
#> [1] 2.287769 2.606098Visualizing the Dynamics Without Process Noise and Measurement Error (n = 5 with Different Initial Condition)
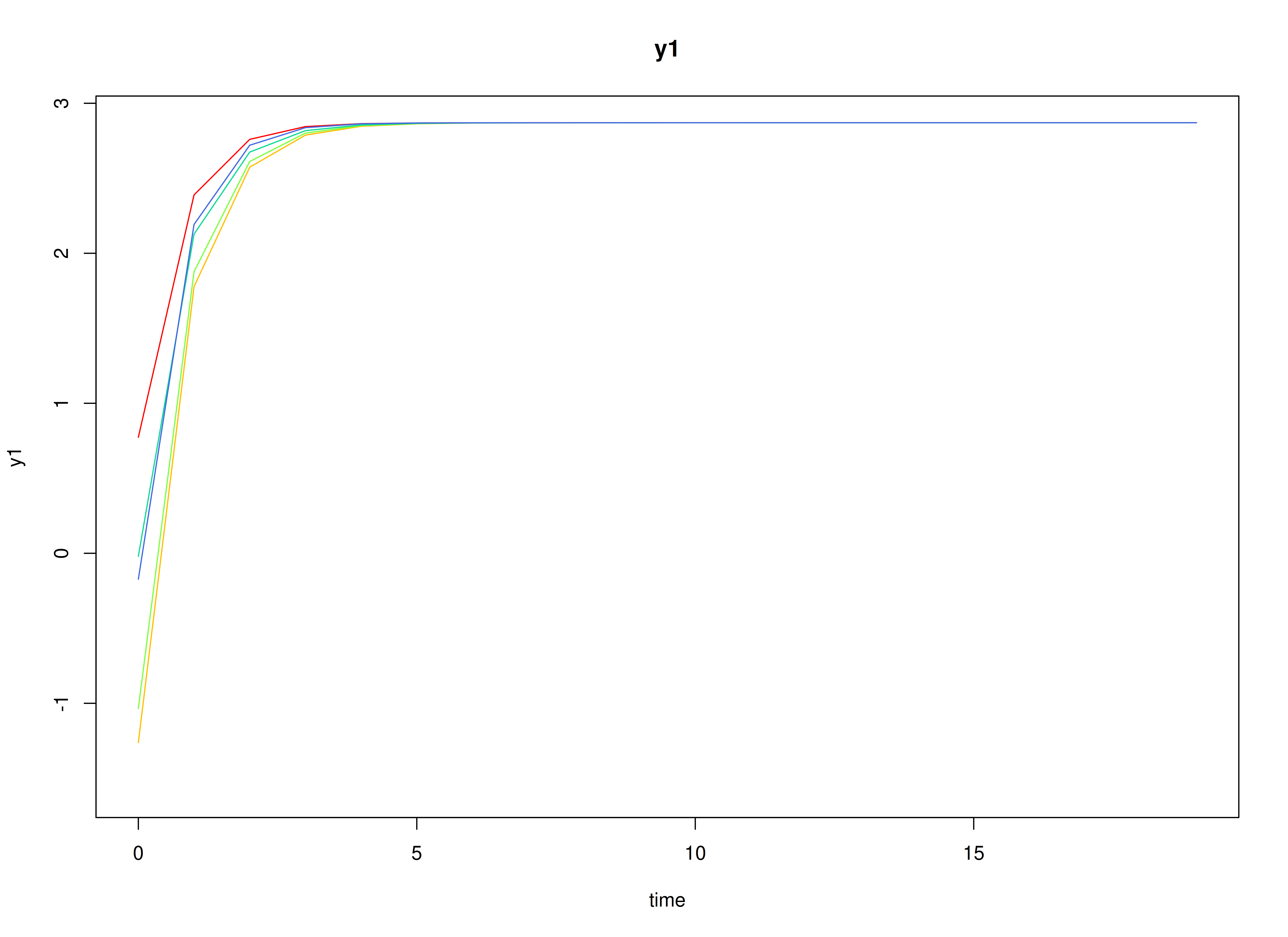
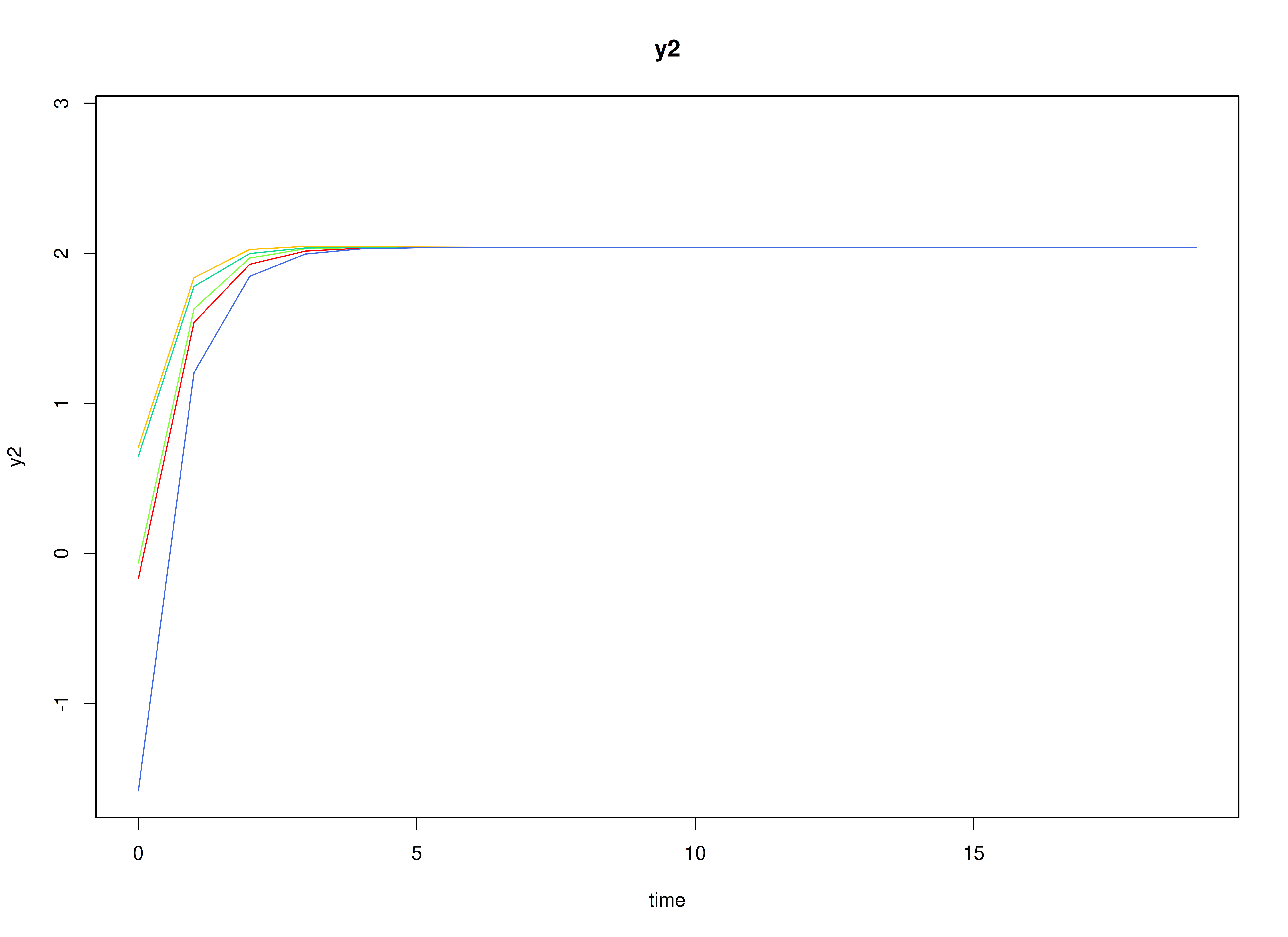
Using the SimSSMVARIVary Function from the
simStateSpace Package to Simulate Data
library(simStateSpace)
sim <- SimSSMVARIVary(
n = n,
time = time,
mu0 = mu0,
sigma0_l = sigma0_l,
alpha = alpha,
beta = beta,
psi_l = psi_l
)
data <- as.data.frame(sim)
head(data)
#> id time y1 y2
#> 1 1 0 2.2494783 1.541912
#> 2 1 1 2.8646254 2.346091
#> 3 1 2 0.8218736 3.348071
#> 4 1 3 2.4900816 2.282659
#> 5 1 4 2.7074414 3.944741
#> 6 1 5 2.9755157 2.667683
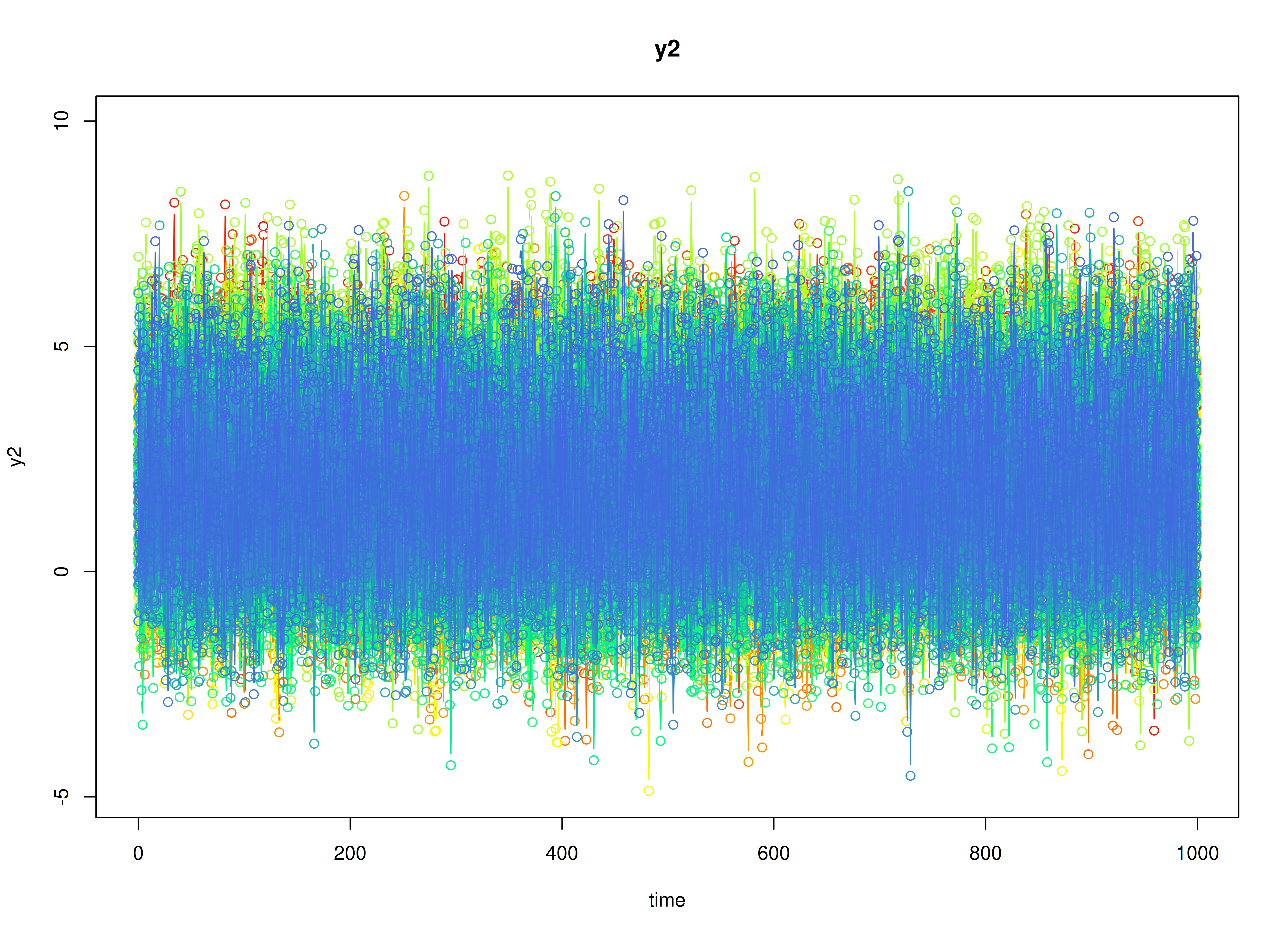
plot(sim)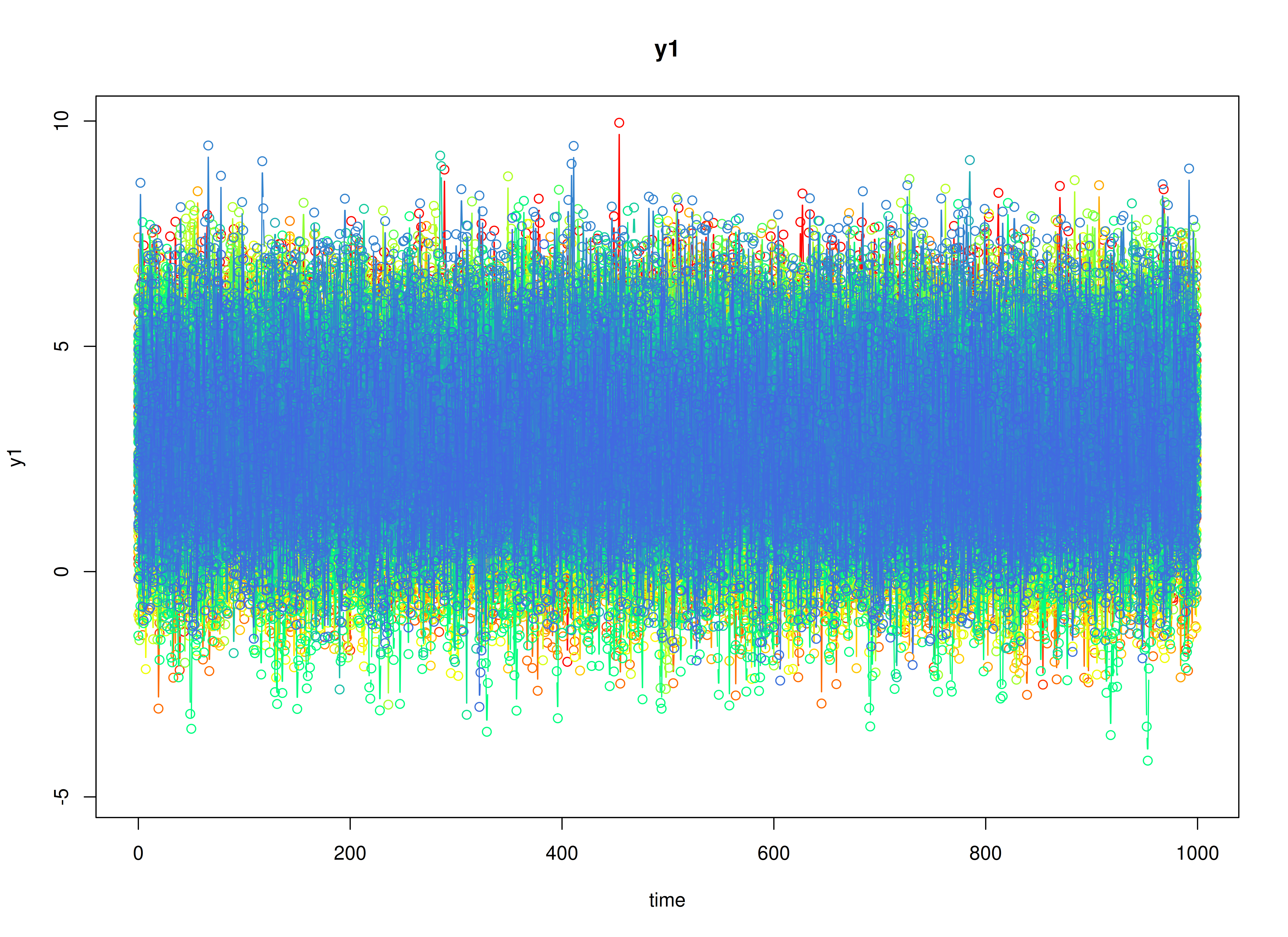
Stage 1: Person-Specific VAR Model
The FitVARMxID function fits a VAR model on each
individual
.
Note: Consider using the argument
ncoresto use multiple cores for parallel processing.
fit <- FitVARMxID(
data = data,
observed = c("y1", "y2"),
id = "id",
center = TRUE,
ncores = parallel::detectCores()
)
summary(
fit,
means = TRUE
)
#> Call:
#> FitVARMxID(data = data, observed = c("y1", "y2"), id = "id",
#> center = TRUE, ncores = parallel::detectCores())
#>
#> Convergence:
#> 100.0%
#>
#> Means of the estimated paramaters per individual.
#> mu[1,1] mu[2,1] beta[1,1] beta[2,1] beta[1,2] beta[2,2] psi[1,1] psi[2,1]
#> 2.9733 2.1842 0.2711 -0.0659 -0.0543 0.2394 1.2932 0.5612
#> psi[2,2]
#> 1.5390Stage 2: Random-Effects Meta-Analysis of Person-Specific Set Points and Dynamics
We synthesize the person-specific estimates to recover population-level effects and their between-person variability. We use a random-effects model so the pooled mean reflects both within-person estimation uncertainty and between-person heterogeneity.
All available parameters are meta-analyzed by default. Setting
cov_dyn = FALSE, meta-analyzes only the set points and
transition matrix. Setting tau_sqr_l_free, such that
covariances between mu and beta are constained
to zero, simplifies the random effects.
tau_sqr_l_free <- matrix(
data = c(
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
TRUE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, FALSE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, FALSE, FALSE, FALSE,
FALSE, FALSE, TRUE, TRUE, FALSE, FALSE,
FALSE, FALSE, TRUE, TRUE, TRUE, FALSE
),
byrow = TRUE,
nrow = 6,
ncol = 6
)
# FALSE values in tau_sqr_l_free will be treated as fixed values.
# TRUE values in tau_sqr_l_free will be treated as starting values.
tau_sqr_l_values <- matrix(
data = 0,
nrow = 6,
ncol = 6
)Note: Consider using the argument
ncoresto use multiple cores for parallel processing.
library(metaDyn)
metavar <- MetaVARMx(
object = fit,
tau_sqr_l_free = tau_sqr_l_free,
tau_sqr_l_values = tau_sqr_l_values,
cov_dyn = FALSE,
ncores = parallel::detectCores()
)
summary(metavar)
#> Call:
#> MetaVARMx(object = fit, tau_sqr_l_free = tau_sqr_l_free, tau_sqr_l_values = tau_sqr_l_values,
#> cov_dyn = FALSE, ncores = parallel::detectCores())
#>
#> Status code:
#> 0
#>
#> Confidence intervals type:
#> Wald
#>
#> Sampling covariance matrix type:
#> Normal
#>
#> est se z p 2.5% 97.5%
#> alpha[1,1] 2.9733 0.1115 26.6781 0.0000 2.7549 3.1918
#> alpha[2,1] 2.1844 0.1052 20.7594 0.0000 1.9781 2.3906
#> alpha[3,1] 0.2720 0.0122 22.3157 0.0000 0.2481 0.2959
#> alpha[4,1] -0.0659 0.0106 -6.2428 0.0000 -0.0866 -0.0452
#> alpha[5,1] -0.0545 0.0047 -11.6738 0.0000 -0.0636 -0.0453
#> alpha[6,1] 0.2401 0.0150 16.0294 0.0000 0.2107 0.2694
#> tau_sqr[1,1] 1.2396 0.1757 7.0558 0.0000 0.8952 1.5839
#> tau_sqr[2,1] 0.5389 0.1291 4.1756 0.0000 0.2859 0.7918
#> tau_sqr[2,2] 1.1040 0.1566 7.0514 0.0000 0.7971 1.4108
#> tau_sqr[3,3] 0.0138 0.0021 6.6038 0.0000 0.0097 0.0179
#> tau_sqr[4,3] 0.0021 0.0013 1.5812 0.1138 -0.0005 0.0046
#> tau_sqr[5,3] 0.0013 0.0006 2.3450 0.0190 0.0002 0.0025
#> tau_sqr[6,3] 0.0037 0.0019 2.0081 0.0446 0.0001 0.0074
#> tau_sqr[4,4] 0.0099 0.0016 6.2656 0.0000 0.0068 0.0130
#> tau_sqr[5,4] 0.0007 0.0005 1.4928 0.1355 -0.0002 0.0017
#> tau_sqr[6,4] 0.0053 0.0017 3.2207 0.0013 0.0021 0.0086
#> tau_sqr[5,5] 0.0013 0.0003 4.2040 0.0000 0.0007 0.0019
#> tau_sqr[6,5] 0.0023 0.0007 3.1303 0.0017 0.0008 0.0037
#> tau_sqr[6,6] 0.0214 0.0032 6.7778 0.0000 0.0152 0.0276
#> i_sqr[1,1] 0.9982 0.0003 3860.4539 0.0000 0.9977 0.9987
#> i_sqr[2,1] 0.9978 0.0003 3176.9990 0.0000 0.9972 0.9984
#> i_sqr[3,1] 0.9293 0.0100 93.3767 0.0000 0.9098 0.9488
#> i_sqr[4,1] 0.8879 0.0159 55.9007 0.0000 0.8568 0.9190
#> i_sqr[5,1] 0.5975 0.0572 10.4451 0.0000 0.4854 0.7096
#> i_sqr[6,1] 0.9537 0.0065 146.4232 0.0000 0.9409 0.9665Normal Theory Confidence Intervals
confint(metavar, level = 0.95)
#> 2.5 % 97.5 %
#> alpha[1,1] 2.754884e+00 3.191767760
#> alpha[2,1] 1.978127e+00 2.390593440
#> alpha[3,1] 2.481423e-01 0.295927294
#> alpha[4,1] -8.662875e-02 -0.045230584
#> alpha[5,1] -6.361191e-02 -0.045322506
#> alpha[6,1] 2.107037e-01 0.269408545
#> tau_sqr[1,1] 8.952303e-01 1.583881949
#> tau_sqr[2,1] 2.859454e-01 0.791842754
#> tau_sqr[2,2] 7.971032e-01 1.410798980
#> tau_sqr[3,3] 9.704755e-03 0.017896751
#> tau_sqr[4,3] -4.944716e-04 0.004622890
#> tau_sqr[5,3] 2.204830e-04 0.002465313
#> tau_sqr[6,3] 8.942248e-05 0.007368261
#> tau_sqr[4,4] 6.796436e-03 0.012984100
#> tau_sqr[5,4] -2.308000e-04 0.001705878
#> tau_sqr[6,4] 2.085974e-03 0.008571963
#> tau_sqr[5,5] 6.881603e-04 0.001890234
#> tau_sqr[6,5] 8.490796e-04 0.003692948
#> tau_sqr[6,6] 1.519136e-02 0.027551438
#> i_sqr[1,1] 9.976655e-01 0.998679034
#> i_sqr[2,1] 9.971649e-01 0.998395992
#> i_sqr[3,1] 9.097638e-01 0.948774247
#> i_sqr[4,1] 8.567713e-01 0.919033710
#> i_sqr[5,1] 4.853568e-01 0.709579238
#> i_sqr[6,1] 9.409422e-01 0.966474135
confint(metavar, level = 0.99)
#> 0.5 % 99.5 %
#> alpha[1,1] 2.6862444124 3.260407197
#> alpha[2,1] 1.9133243881 2.455396538
#> alpha[3,1] 0.2406346851 0.303434868
#> alpha[4,1] -0.0931328751 -0.038726460
#> alpha[5,1] -0.0664853861 -0.042449032
#> alpha[6,1] 0.2014804830 0.278631747
#> tau_sqr[1,1] 0.7870352302 1.692076971
#> tau_sqr[2,1] 0.2064631929 0.871324985
#> tau_sqr[2,2] 0.7006846514 1.507217569
#> tau_sqr[3,3] 0.0084176997 0.019183807
#> tau_sqr[4,3] -0.0012984674 0.005426885
#> tau_sqr[5,3] -0.0001322053 0.002818001
#> tau_sqr[6,3] -0.0010541659 0.008511849
#> tau_sqr[4,4] 0.0058242832 0.013956253
#> tau_sqr[5,4] -0.0005350741 0.002010152
#> tau_sqr[6,4] 0.0010669519 0.009590985
#> tau_sqr[5,5] 0.0004993009 0.002079093
#> tau_sqr[6,5] 0.0004022755 0.004139752
#> tau_sqr[6,6] 0.0132494547 0.029493346
#> i_sqr[1,1] 0.9975062441 0.998838275
#> i_sqr[2,1] 0.9969714633 0.998589413
#> i_sqr[3,1] 0.9036348181 0.954903232
#> i_sqr[4,1] 0.8469891943 0.928815839
#> i_sqr[5,1] 0.4501289257 0.744807133
#> i_sqr[6,1] 0.9369308440 0.970485493Robust Confidence Intervals
confint(metavar, level = 0.95, robust = TRUE)
#> 2.5 % 97.5 %
#> alpha[1,1] 2.7537802250 3.192871385
#> alpha[2,1] 1.9770921922 2.391628734
#> alpha[3,1] 0.2480754679 0.295994085
#> alpha[4,1] -0.0867440021 -0.045115333
#> alpha[5,1] -0.0636814145 -0.045253003
#> alpha[6,1] 0.2105511450 0.269561085
#> tau_sqr[1,1] 0.9013903464 1.577721855
#> tau_sqr[2,1] 0.2794401039 0.798348074
#> tau_sqr[2,2] 0.8278224407 1.380079780
#> tau_sqr[3,3] 0.0091150505 0.018486456
#> tau_sqr[4,3] -0.0002357785 0.004364197
#> tau_sqr[5,3] 0.0003160409 0.002369755
#> tau_sqr[6,3] -0.0005935103 0.008051193
#> tau_sqr[4,4] 0.0072591008 0.012521436
#> tau_sqr[5,4] -0.0001960227 0.001671100
#> tau_sqr[6,4] 0.0023412838 0.008316653
#> tau_sqr[5,5] 0.0007204276 0.001857966
#> tau_sqr[6,5] 0.0011175131 0.003424514
#> tau_sqr[6,6] 0.0134983999 0.029244401
#> i_sqr[1,1] 0.9976745507 0.998669968
#> i_sqr[2,1] 0.9972265088 0.998334367
#> i_sqr[3,1] 0.9069556163 0.951582434
#> i_sqr[4,1] 0.8614268211 0.914378213
#> i_sqr[5,1] 0.4913757870 0.703560271
#> i_sqr[6,1] 0.9374452033 0.969971134
confint(metavar, level = 0.99, robust = TRUE)
#> 0.5 % 99.5 %
#> alpha[1,1] 2.684794e+00 3.261857606
#> alpha[2,1] 1.911964e+00 2.456757145
#> alpha[3,1] 2.405469e-01 0.303522645
#> alpha[4,1] -9.328434e-02 -0.038574995
#> alpha[5,1] -6.657673e-02 -0.042357690
#> alpha[6,1] 2.012800e-01 0.278832219
#> tau_sqr[1,1] 7.951310e-01 1.683981237
#> tau_sqr[2,1] 1.979138e-01 0.879874425
#> tau_sqr[2,2] 7.410565e-01 1.466845696
#> tau_sqr[3,3] 7.642696e-03 0.019958811
#> tau_sqr[4,3] -9.584870e-04 0.005086905
#> tau_sqr[5,3] -6.620909e-06 0.002692416
#> tau_sqr[6,3] -1.951692e-03 0.009409375
#> tau_sqr[4,4] 6.432328e-03 0.013348208
#> tau_sqr[5,4] -4.893690e-04 0.001964447
#> tau_sqr[6,4] 1.402485e-03 0.009255452
#> tau_sqr[5,5] 5.417073e-04 0.002036687
#> tau_sqr[6,5] 7.550569e-04 0.003786971
#> tau_sqr[6,6] 1.102452e-02 0.031718277
#> i_sqr[1,1] 9.975182e-01 0.998826359
#> i_sqr[2,1] 9.970525e-01 0.998508425
#> i_sqr[3,1] 8.999442e-01 0.958593815
#> i_sqr[4,1] 8.531076e-01 0.922697479
#> i_sqr[5,1] 4.580392e-01 0.736896870
#> i_sqr[6,1] 9.323350e-01 0.975081328- The fixed part of the random-effects model gives pooled means .
- The random part yields between-person covariances () quantifying heterogeneity in set point () and dynamics () across individuals.
means <- extract(metavar, what = "alpha")
means
#> alpha
#> y1 2.97332580
#> y2 2.18436046
#> y3 0.27203478
#> y4 -0.06592967
#> y5 -0.05446721
#> y6 0.24005612
covariances <- extract(metavar, what = "tau_sqr")
covariances
#> y1 y2 y3 y4 y5 y6
#> y1 1.2395561 0.5388941 0.000000000 0.0000000000 0.0000000000 0.000000000
#> y2 0.5388941 1.1039511 0.000000000 0.0000000000 0.0000000000 0.000000000
#> y3 0.0000000 0.0000000 0.013800753 0.0020642090 0.0013428978 0.003728842
#> y4 0.0000000 0.0000000 0.002064209 0.0098902681 0.0007375388 0.005328969
#> y5 0.0000000 0.0000000 0.001342898 0.0007375388 0.0012891970 0.002271014
#> y6 0.0000000 0.0000000 0.003728842 0.0053289686 0.0022710137 0.021371400Summary
This vignette demonstrates a two-stage hierarchical estimation
approach for dynamic systems: 1. individual-level DT-VAR estimation,
and
2. population-level meta-analysis of person-specific set points and
dynamics.