Discrete-Time Vector Autoregressive Model (center = TRUE)
Ivan Jacob Agaloos Pesigan
2026-06-08
Source:vignettes/dt-center-true.Rmd
dt-center-true.RmdModel
The measurement model is given by where , and are random variables.
The dynamic structure is given by where , , and are random variables, and , , and are model parameters. Here, is a vector of latent variables at time and individual , represents a vector of latent variables at time and individual , and represents a vector of dynamic noise at time and individual . denotes a vector of set-points, a matrix of autoregression and cross regression coefficients, and the covariance matrix of .
An alternative representation of the dynamic noise is given by where .
Data Generation
Notation
Let be the number of time points and be the number of individuals.
Let the initial condition be given by
Let the set-point vector be given by
Let the transition matrix be given by
Let the dynamic process noise be given by
R Function Arguments
n
#> [1] 100
time
#> [1] 1000
mu0
#> [1] 3.049353 6.154380 4.885913
sigma0
#> [,1] [,2] [,3]
#> [1,] 0.19607843 0.1183232 0.02985385
#> [2,] 0.11832319 0.3437711 0.13818551
#> [3,] 0.02985385 0.1381855 0.26638284
sigma0_l # sigma0_l <- t(chol(sigma0))
#> [,1] [,2] [,3]
#> [1,] 0.44280744 0.0000000 0.000000
#> [2,] 0.26721139 0.5218900 0.000000
#> [3,] 0.06741949 0.2302597 0.456966
alpha
#> [1] 0.9148060 0.9370754 0.2861395
beta
#> [,1] [,2] [,3]
#> [1,] 0.7 0.0 0.0
#> [2,] 0.5 0.6 0.0
#> [3,] -0.1 0.4 0.5
psi
#> [,1] [,2] [,3]
#> [1,] 0.1 0.0 0.0
#> [2,] 0.0 0.1 0.0
#> [3,] 0.0 0.0 0.1
psi_l # psi_l <- t(chol(psi))
#> [,1] [,2] [,3]
#> [1,] 0.3162278 0.0000000 0.0000000
#> [2,] 0.0000000 0.3162278 0.0000000
#> [3,] 0.0000000 0.0000000 0.3162278
# set-point
mu
#> [1] 3.049353 6.154380 4.885913Using the SimSSMVARFixed Function from the
simStateSpace Package to Simulate Data
library(simStateSpace)
sim <- SimSSMVARFixed(
n = n,
time = time,
mu0 = mu0,
sigma0_l = sigma0_l,
alpha = alpha,
beta = beta,
psi_l = psi_l
)
data <- as.data.frame(sim)
head(data)
#> id time y1 y2 y3
#> 1 1 0 3.375463 5.964917 3.883475
#> 2 1 1 3.136727 5.909374 3.866987
#> 3 1 2 3.215198 6.462812 3.995227
#> 4 1 3 3.427307 6.408953 4.408847
#> 5 1 4 3.108728 6.717274 5.303110
#> 6 1 5 3.885159 7.141896 5.024148
summary(data)
#> id time y1 y2
#> Min. : 1.00 Min. : 0.0 Min. :1.169 Min. :3.552
#> 1st Qu.: 25.75 1st Qu.:249.8 1st Qu.:2.755 1st Qu.:5.767
#> Median : 50.50 Median :499.5 Median :3.054 Median :6.160
#> Mean : 50.50 Mean :499.5 Mean :3.055 Mean :6.163
#> 3rd Qu.: 75.25 3rd Qu.:749.2 3rd Qu.:3.356 3rd Qu.:6.558
#> Max. :100.00 Max. :999.0 Max. :5.029 Max. :8.619
#> y3
#> Min. :2.460
#> 1st Qu.:4.544
#> Median :4.893
#> Mean :4.895
#> 3rd Qu.:5.245
#> Max. :7.672
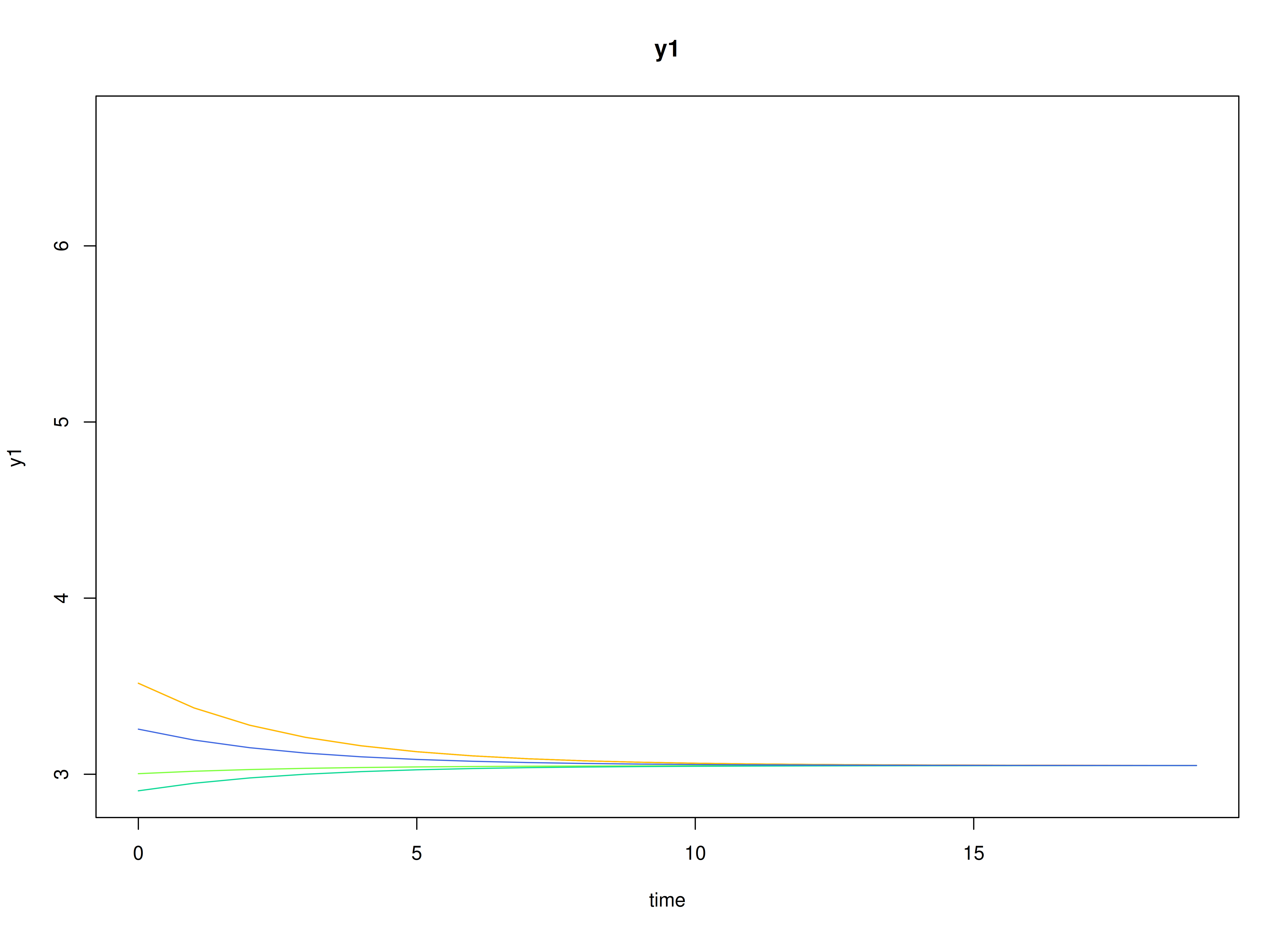
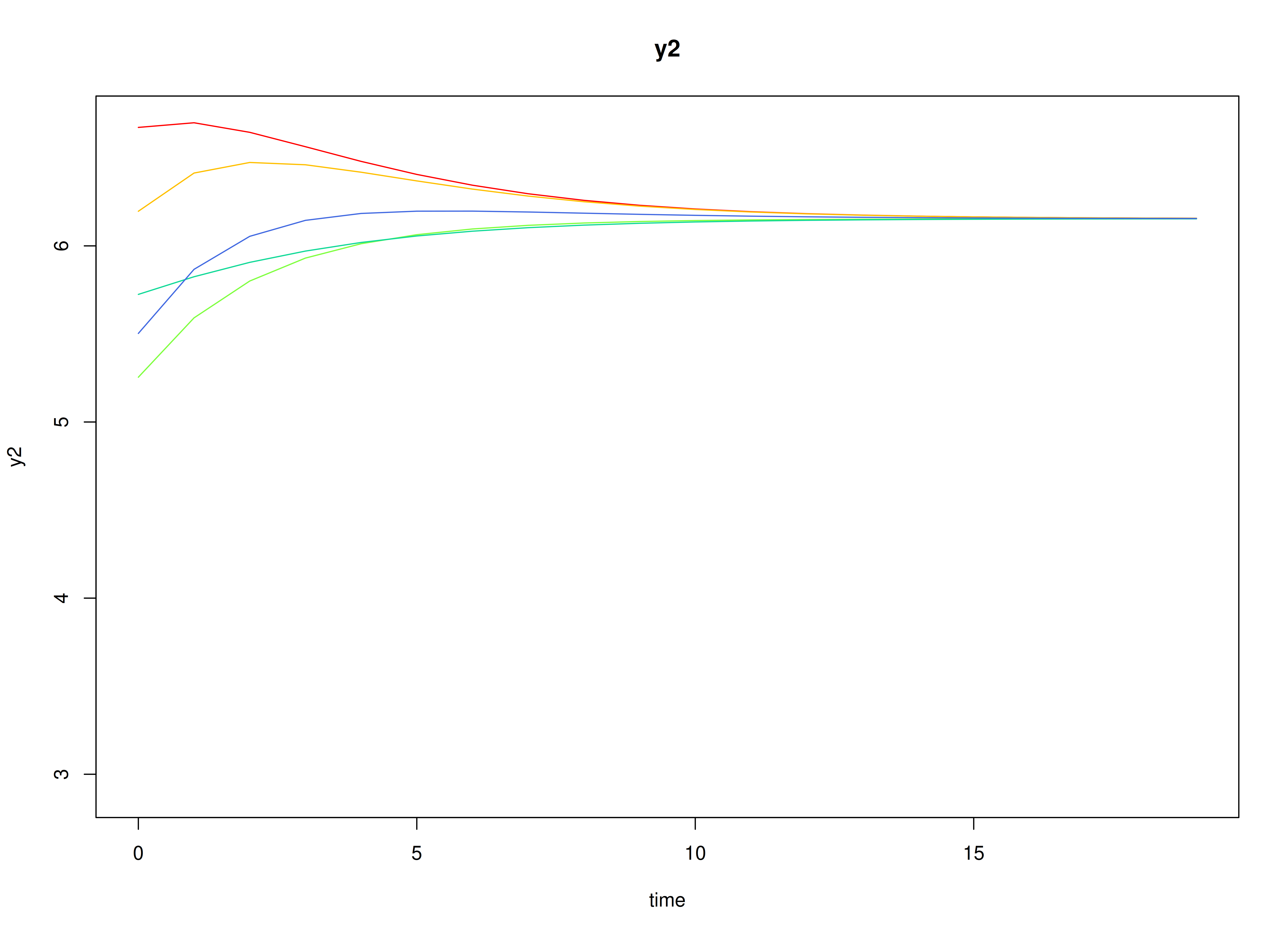
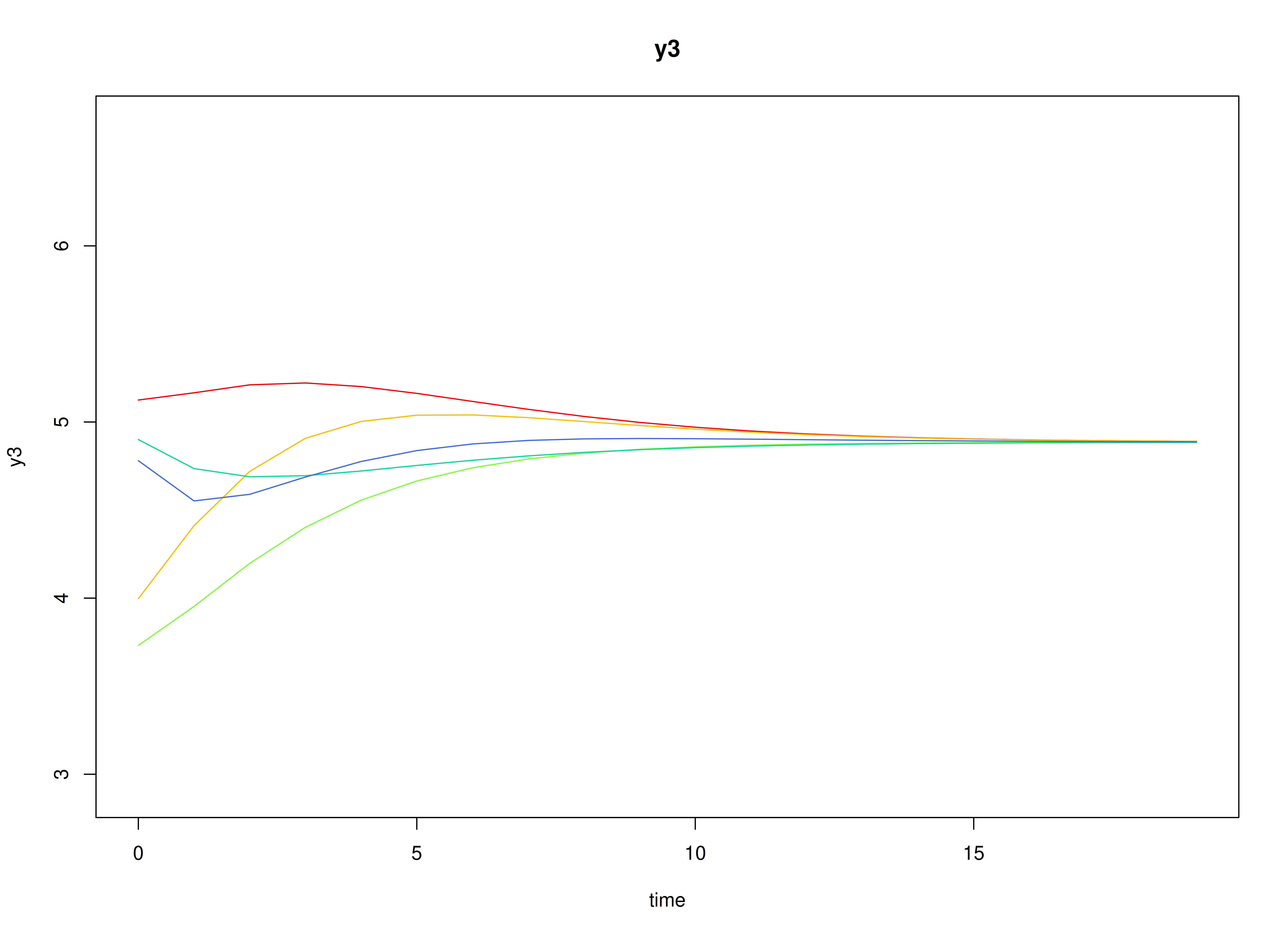
plot(sim)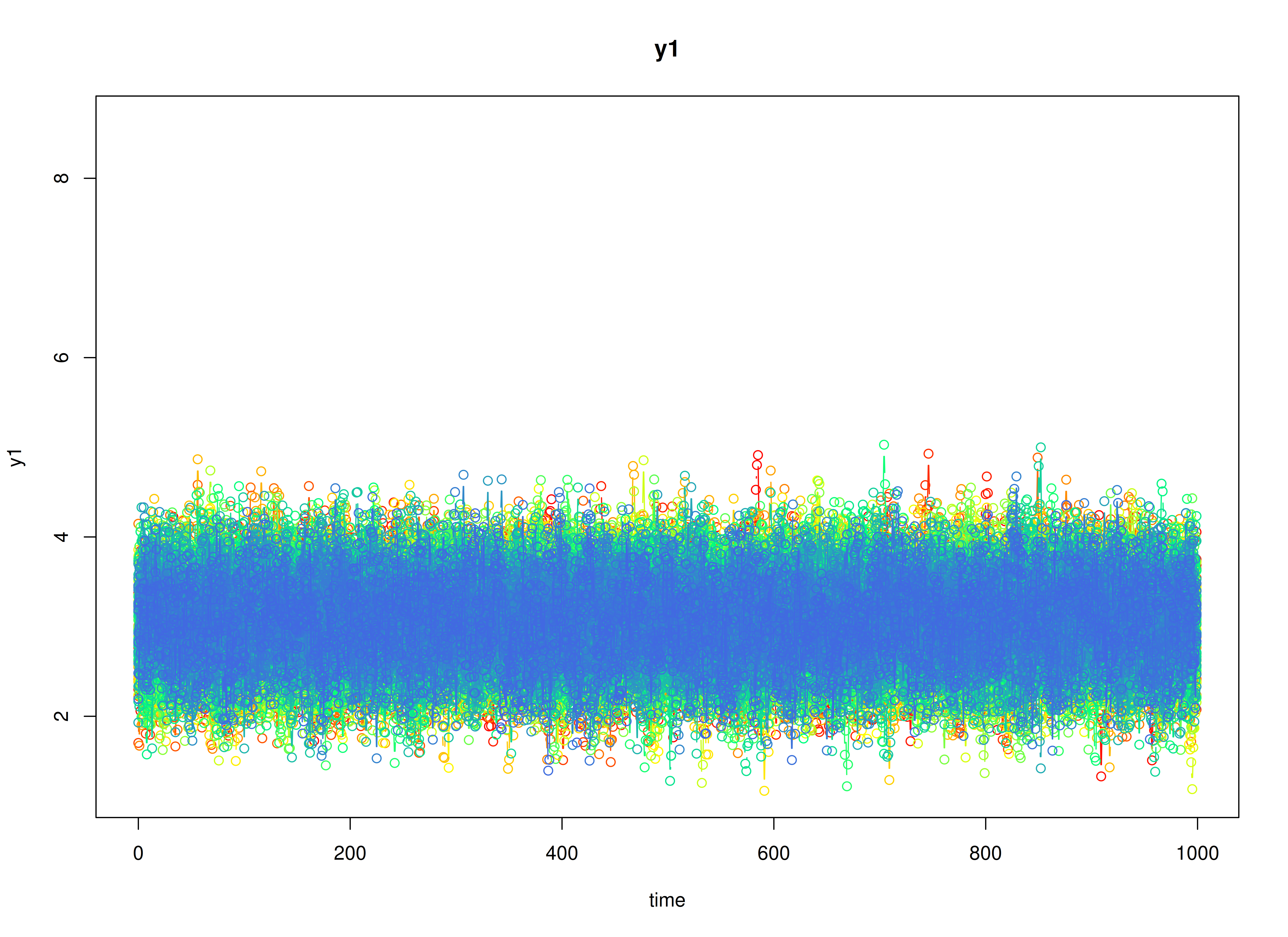
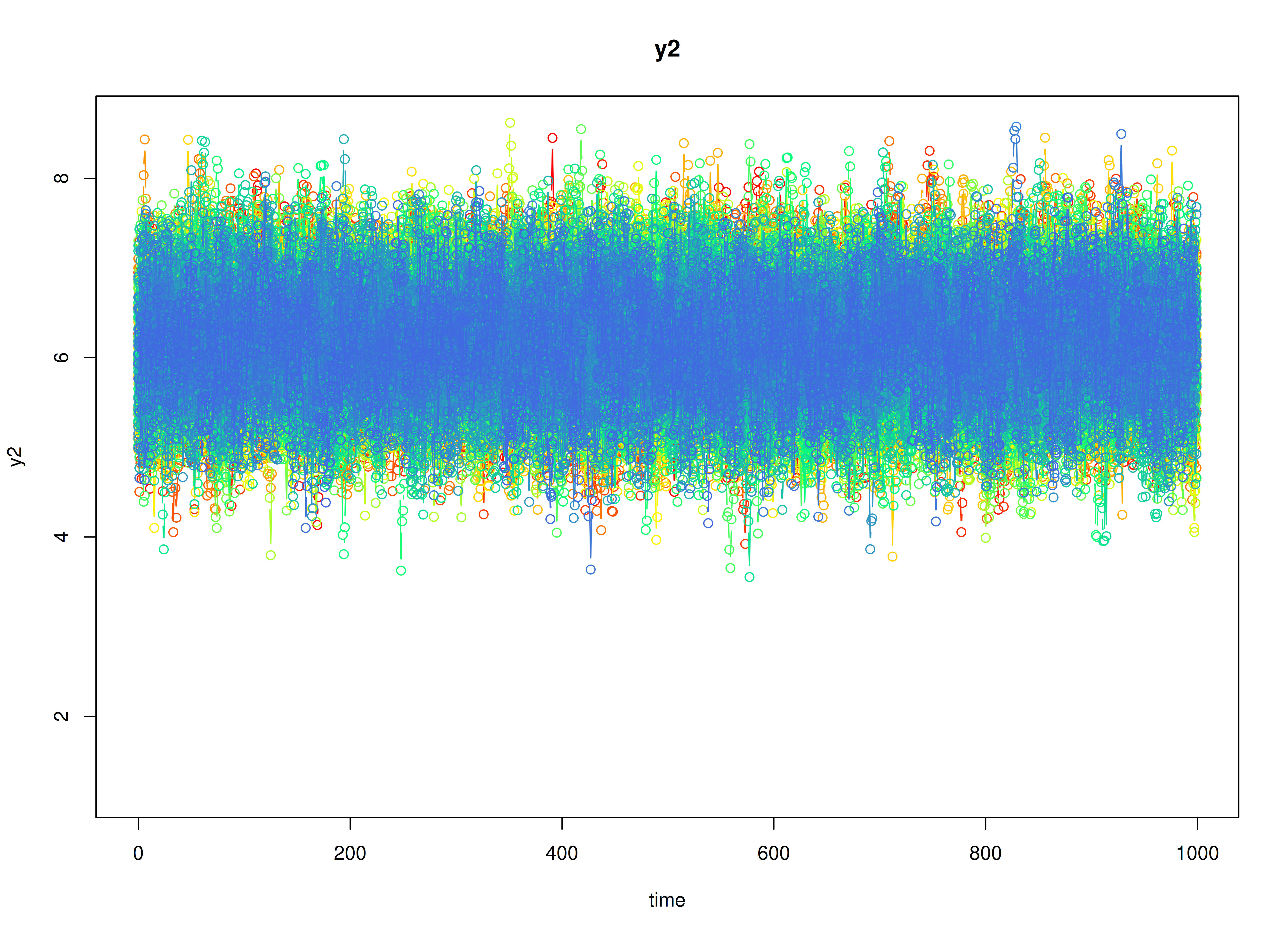
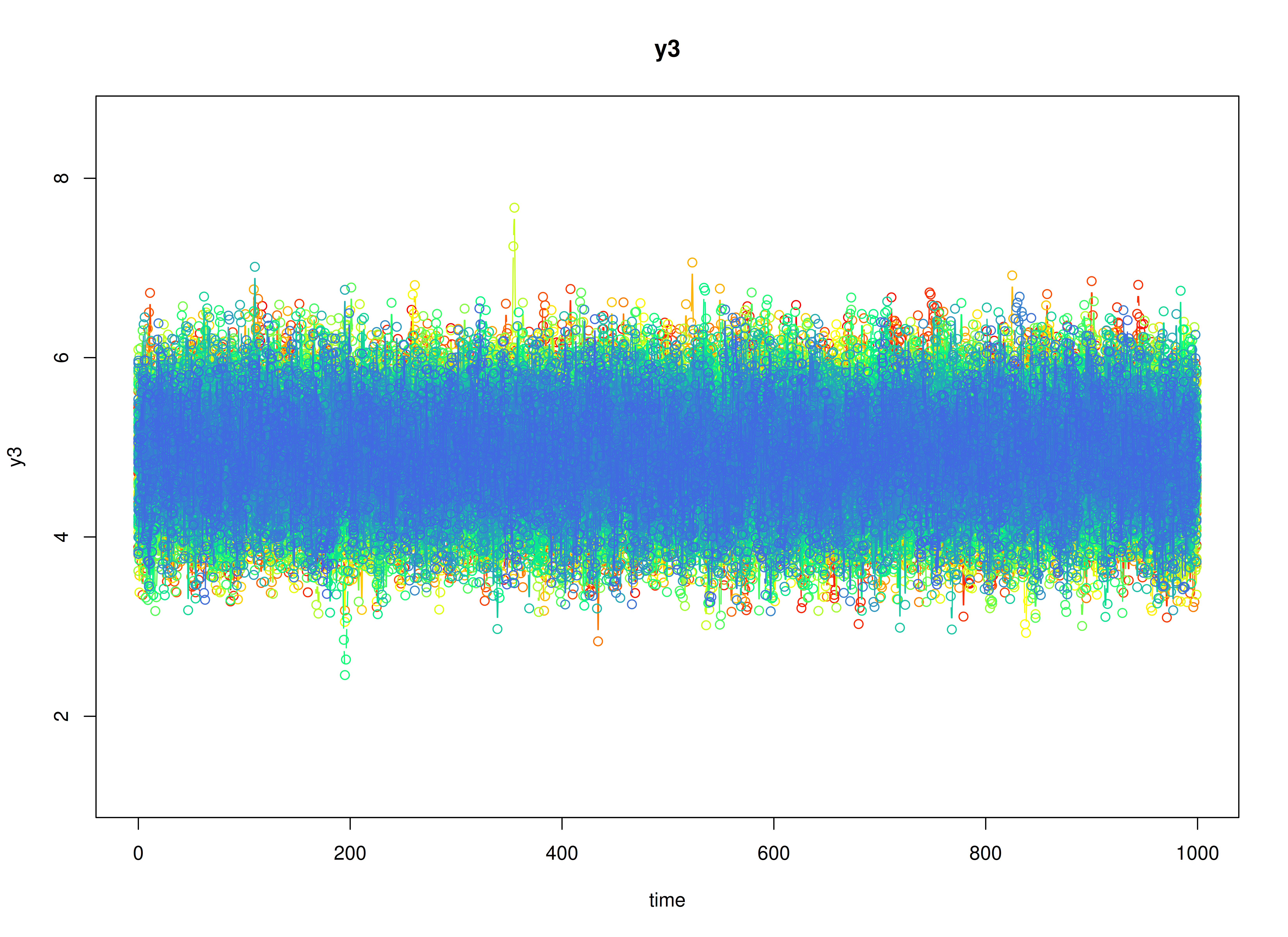
Model Fitting
The FitVARMxID function fits a DT-VAR model on each
individual
.
Note: Consider using the argument
ncoresto use multiple cores for parallel processing.
fit <- FitVARMxID(
data = data,
observed = c("y1", "y2", "y3"),
id = "id",
center = TRUE,
ncores = parallel::detectCores()
)Parameter estimates
summary(
fit,
means = TRUE,
ncores = parallel::detectCores()
)
#> Call:
#> FitVARMxID(data = data, observed = c("y1", "y2", "y3"), id = "id",
#> center = TRUE, ncores = parallel::detectCores())
#>
#> Convergence:
#> 100.0%
#>
#> Means of the estimated paramaters per individual.
#> mu[1,1] mu[2,1] mu[3,1] beta[1,1] beta[2,1] beta[3,1] beta[1,2] beta[2,2]
#> 3.0548 6.1622 4.8943 0.7001 0.4990 -0.0991 -0.0022 0.5998
#> beta[3,2] beta[1,3] beta[2,3] beta[3,3] psi[1,1] psi[2,1] psi[3,1] psi[2,2]
#> 0.4038 -0.0051 -0.0022 0.4982 0.0997 0.0004 -0.0003 0.0990
#> psi[3,2] psi[3,3]
#> 0.0005 0.0994