Discrete-Time Vector Autoregressive Model (center = FALSE)
Ivan Jacob Agaloos Pesigan
2026-06-08
Source:vignettes/dt-center-false.Rmd
dt-center-false.RmdModel
The measurement model is given by where , and are random variables.
The dynamic structure is given by where , , and are random variables, and , , and are model parameters. Here, is a vector of latent variables at time and individual , represents a vector of latent variables at time and individual , and represents a vector of dynamic noise at time and individual . denotes a vector of intercepts, a matrix of autoregression and cross regression coefficients, and the covariance matrix of .
An alternative representation of the dynamic noise is given by where .
Data Generation
Notation
Let be the number of time points and be the number of individuals.
Let the initial condition be given by
Let the constant vector be given by
Let the transition matrix be given by
Let the dynamic process noise be given by
R Function Arguments
n
#> [1] 100
time
#> [1] 1000
mu0
#> [1] 3.049353 6.154380 4.885913
sigma0
#> [,1] [,2] [,3]
#> [1,] 0.19607843 0.1183232 0.02985385
#> [2,] 0.11832319 0.3437711 0.13818551
#> [3,] 0.02985385 0.1381855 0.26638284
sigma0_l # sigma0_l <- t(chol(sigma0))
#> [,1] [,2] [,3]
#> [1,] 0.44280744 0.0000000 0.000000
#> [2,] 0.26721139 0.5218900 0.000000
#> [3,] 0.06741949 0.2302597 0.456966
alpha
#> [1] 0.9148060 0.9370754 0.2861395
beta
#> [,1] [,2] [,3]
#> [1,] 0.7 0.0 0.0
#> [2,] 0.5 0.6 0.0
#> [3,] -0.1 0.4 0.5
psi
#> [,1] [,2] [,3]
#> [1,] 0.1 0.0 0.0
#> [2,] 0.0 0.1 0.0
#> [3,] 0.0 0.0 0.1
psi_l # psi_l <- t(chol(psi))
#> [,1] [,2] [,3]
#> [1,] 0.3162278 0.0000000 0.0000000
#> [2,] 0.0000000 0.3162278 0.0000000
#> [3,] 0.0000000 0.0000000 0.3162278
# set-point
mu
#> [1] 3.049353 6.154380 4.885913Using the SimSSMVARFixed Function from the
simStateSpace Package to Simulate Data
library(simStateSpace)
sim <- SimSSMVARFixed(
n = n,
time = time,
mu0 = mu0,
sigma0_l = sigma0_l,
alpha = alpha,
beta = beta,
psi_l = psi_l
)
data <- as.data.frame(sim)
head(data)
#> id time y1 y2 y3
#> 1 1 0 3.375463 5.964917 3.883475
#> 2 1 1 3.136727 5.909374 3.866987
#> 3 1 2 3.215198 6.462812 3.995227
#> 4 1 3 3.427307 6.408953 4.408847
#> 5 1 4 3.108728 6.717274 5.303110
#> 6 1 5 3.885159 7.141896 5.024148
summary(data)
#> id time y1 y2
#> Min. : 1.00 Min. : 0.0 Min. :1.169 Min. :3.552
#> 1st Qu.: 25.75 1st Qu.:249.8 1st Qu.:2.755 1st Qu.:5.767
#> Median : 50.50 Median :499.5 Median :3.054 Median :6.160
#> Mean : 50.50 Mean :499.5 Mean :3.055 Mean :6.163
#> 3rd Qu.: 75.25 3rd Qu.:749.2 3rd Qu.:3.356 3rd Qu.:6.558
#> Max. :100.00 Max. :999.0 Max. :5.029 Max. :8.619
#> y3
#> Min. :2.460
#> 1st Qu.:4.544
#> Median :4.893
#> Mean :4.895
#> 3rd Qu.:5.245
#> Max. :7.672
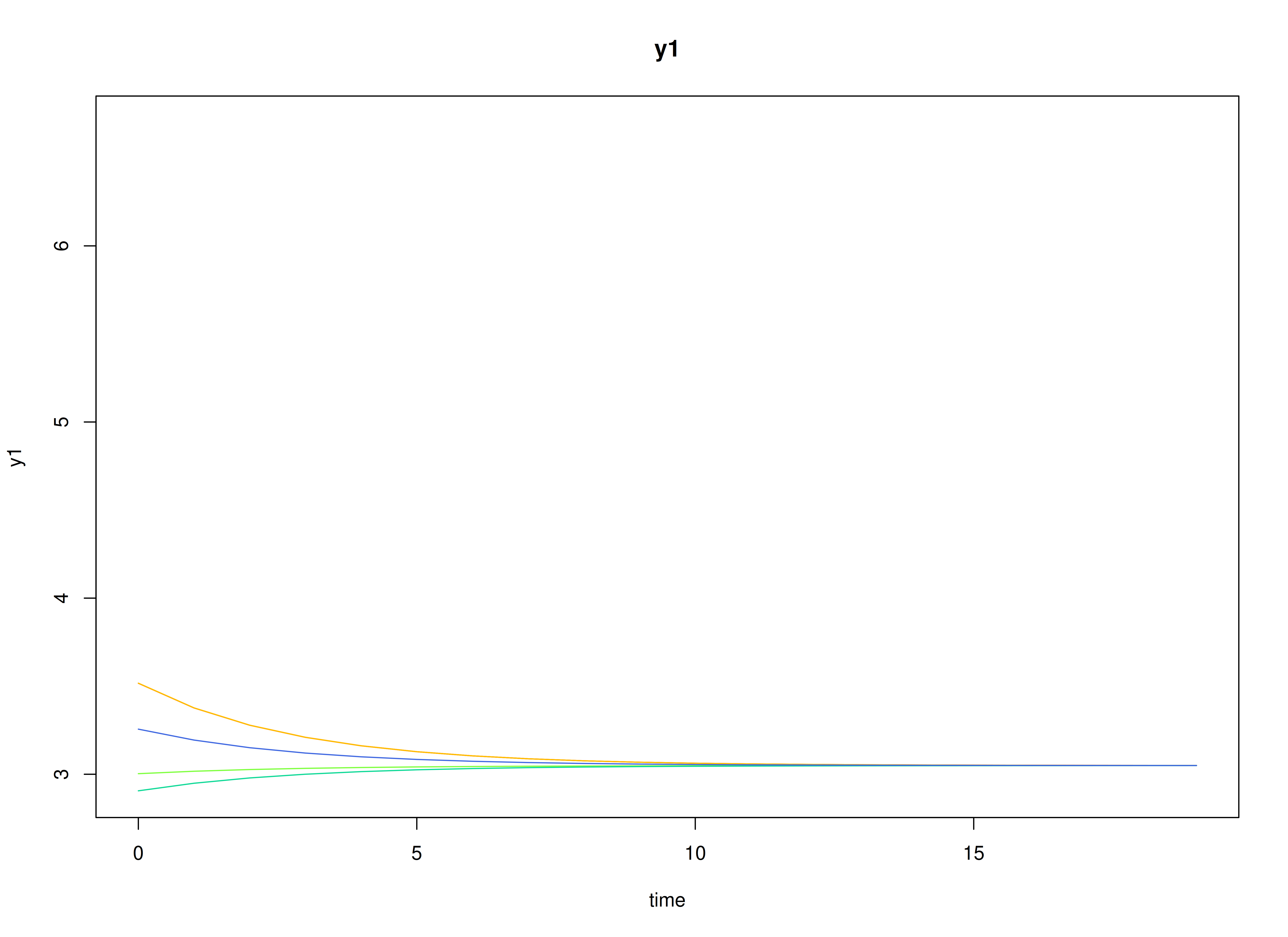
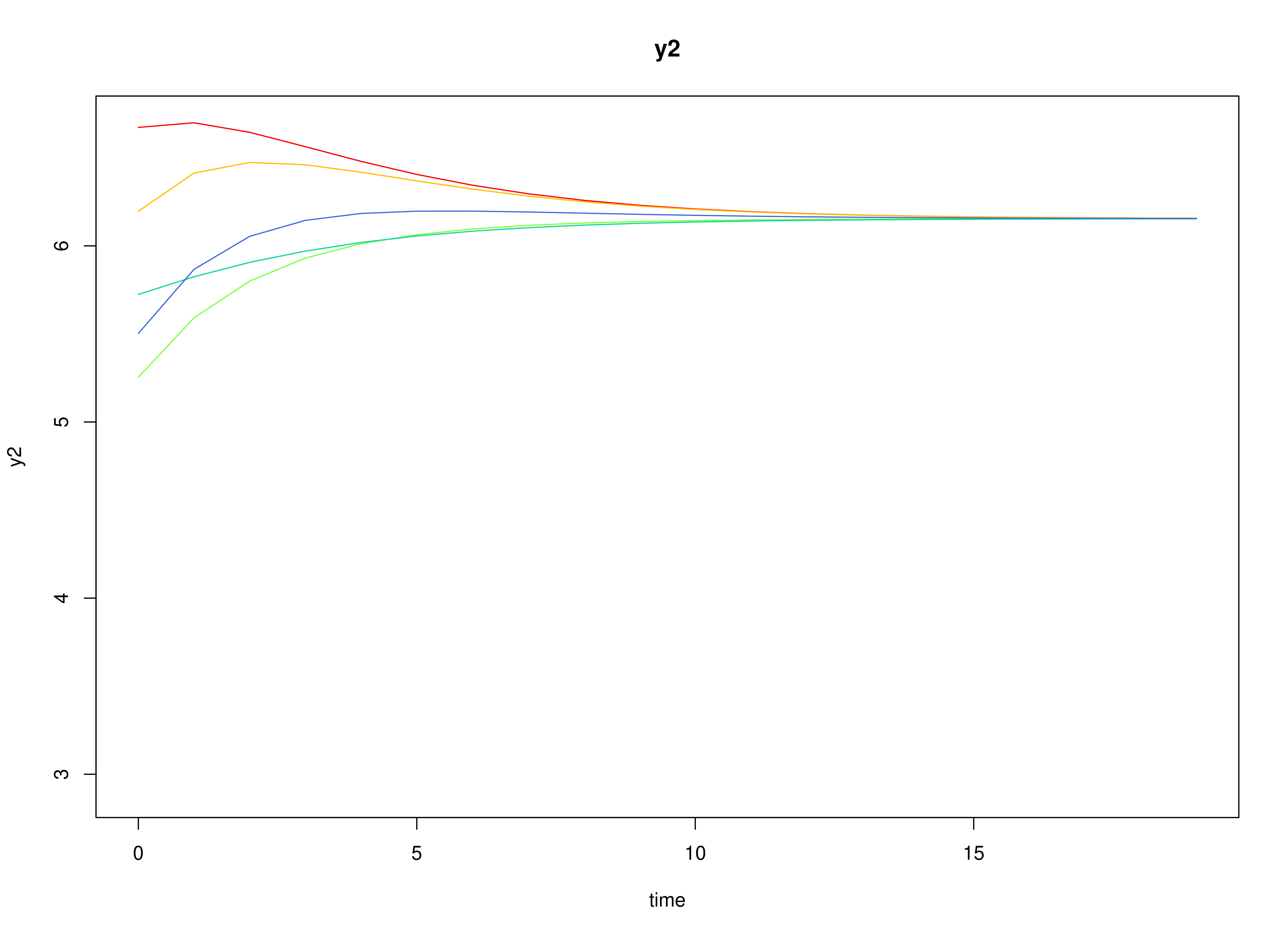
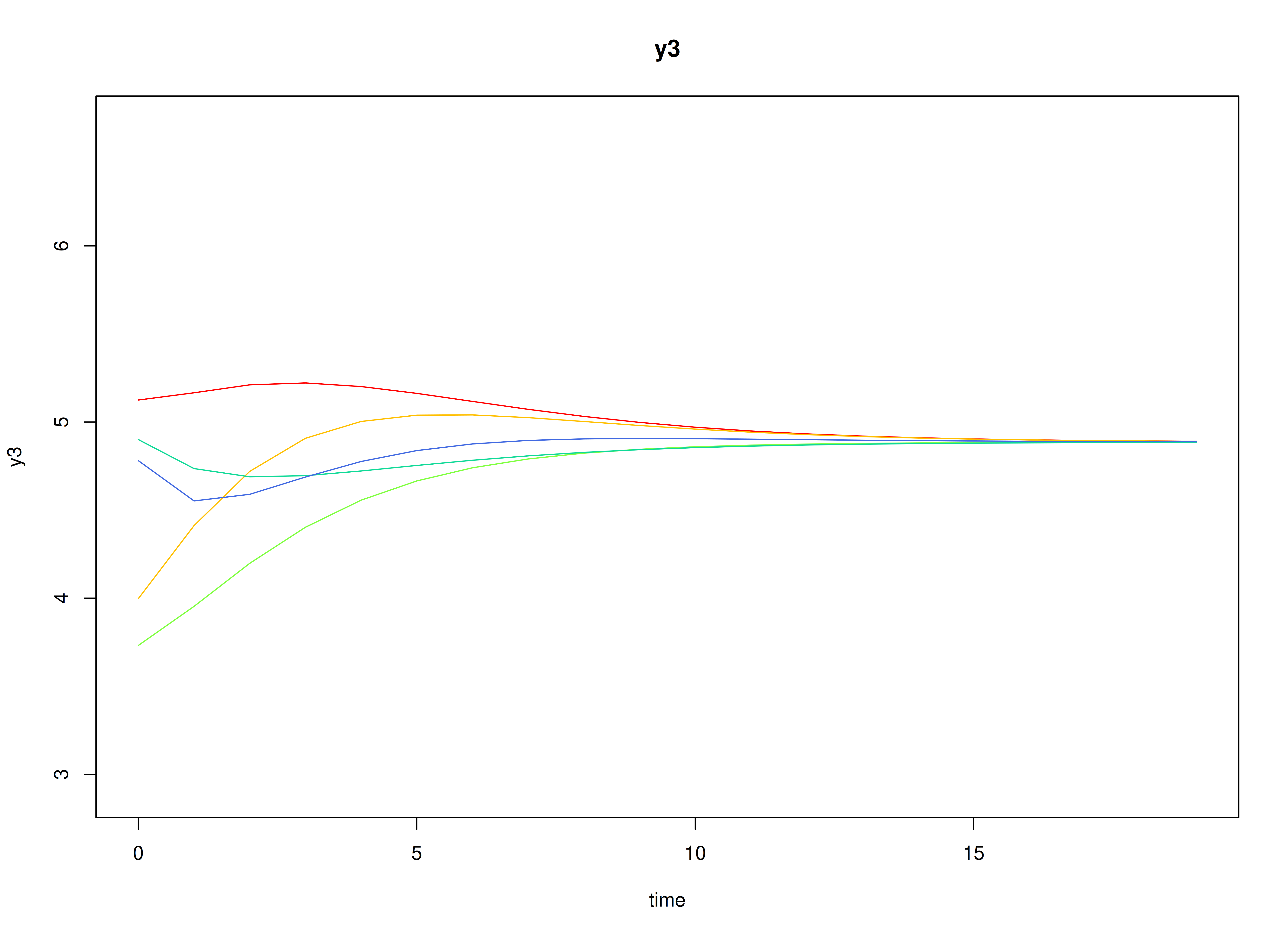
plot(sim)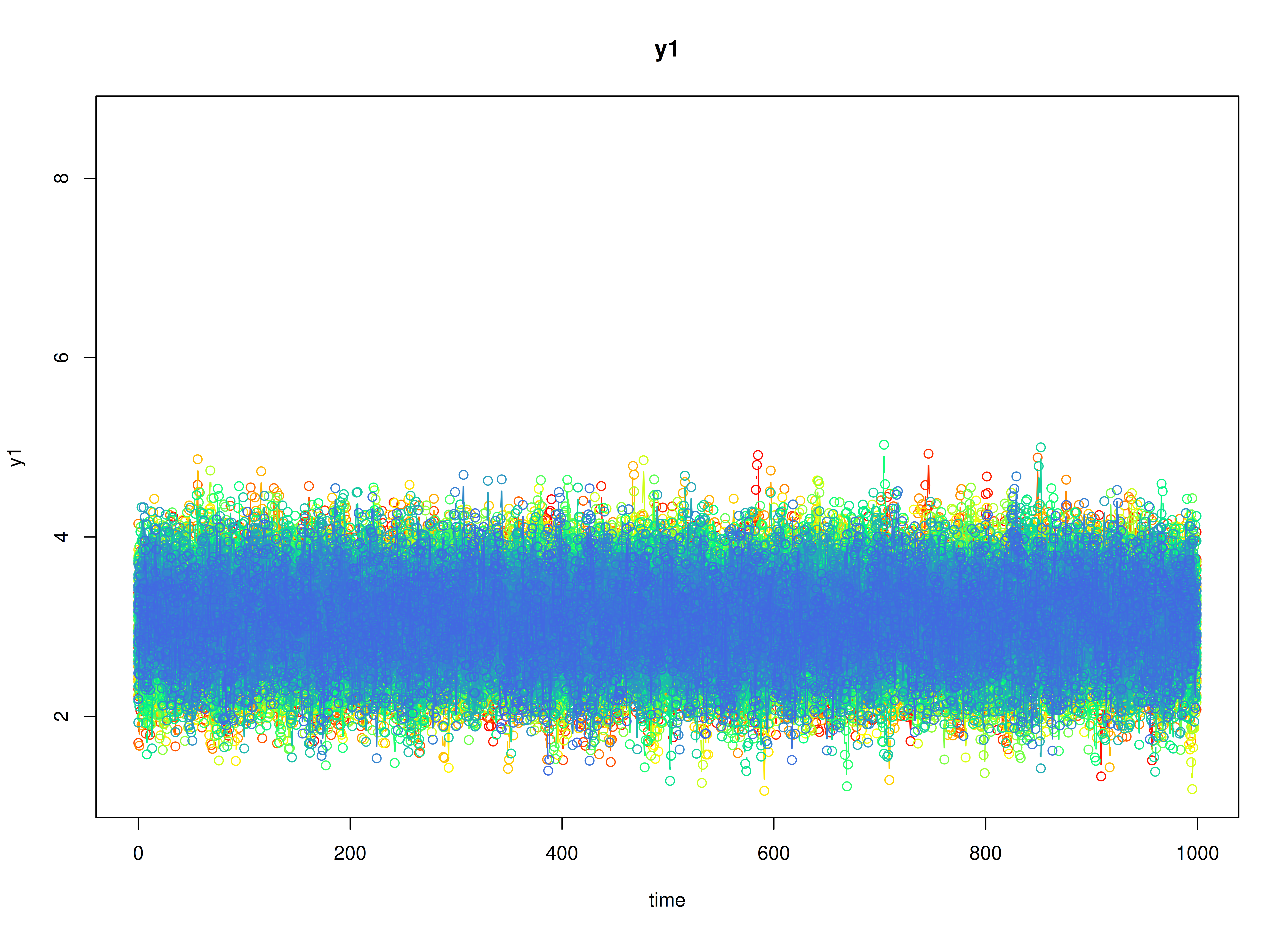
Model Fitting
The FitVARMxID function fits a DT-VAR model on each
individual
.
Note: Consider using the argument
ncoresto use multiple cores for parallel processing.
fit <- FitVARMxID(
data = data,
observed = c("y1", "y2", "y3"),
id = "id",
center = FALSE,
ncores = parallel::detectCores()
)Parameter estimates
summary(
fit,
means = TRUE,
ncores = parallel::detectCores()
)
#> Call:
#> FitVARMxID(data = data, observed = c("y1", "y2", "y3"), id = "id",
#> center = FALSE, ncores = parallel::detectCores())
#>
#> Convergence:
#> 100.0%
#>
#> Means of the estimated paramaters per individual.
#> alpha[1,1] alpha[2,1] alpha[3,1] beta[1,1] beta[2,1] beta[3,1] beta[1,2]
#> 0.9544 0.9524 0.2701 0.7001 0.4990 -0.0991 -0.0022
#> beta[2,2] beta[3,2] beta[1,3] beta[2,3] beta[3,3] psi[1,1] psi[2,1]
#> 0.5998 0.4038 -0.0051 -0.0022 0.4982 0.0997 0.0004
#> psi[3,1] psi[2,2] psi[3,2] psi[3,3]
#> -0.0003 0.0990 0.0005 0.0994